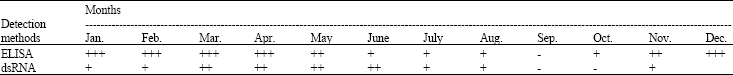Research Article
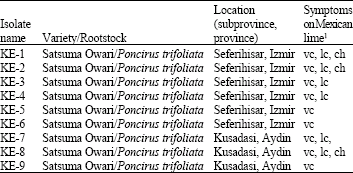
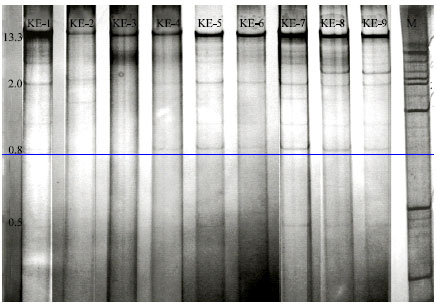
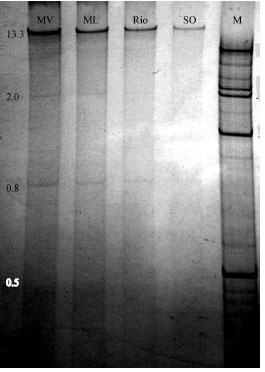
Investigation of Biological Properties, Double-Stranded RNA Patterns and Antigen Concentration in Satsuma Owari Mandarins Infected with Citrus tristeza virus in Aegean Region of Turkey
Department of Plant Protection, Faculty of Agriculture, Canakkale Onsekiz Mart University, 17020, Canakkale, Turkey
Serkan Onder
Department of Plant Protection, Faculty of Agriculture, Canakkale Onsekiz Mart University, 17020, Canakkale, Turkey
Mustafa Gumus
Department of Plant Protection, Faculty of Agriculture, Ege University, 35100, Bornova, Izmir, Turkey