Research Article
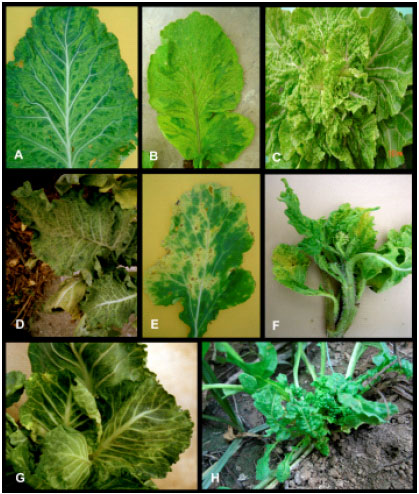
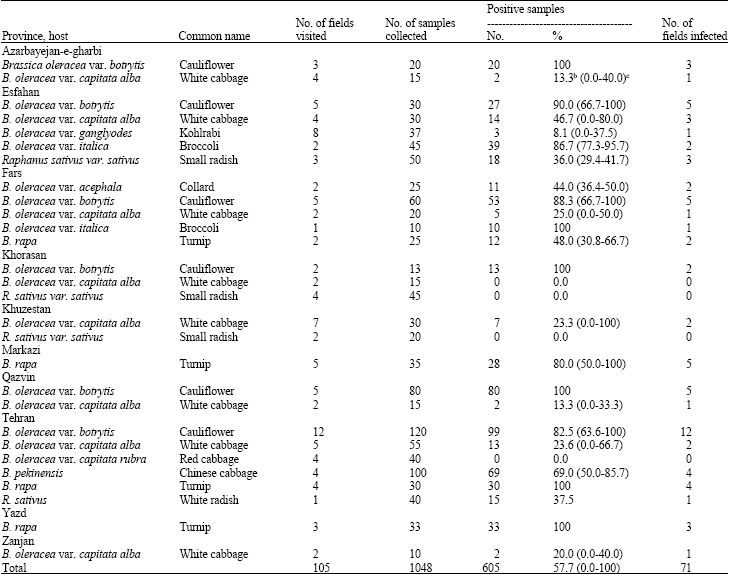
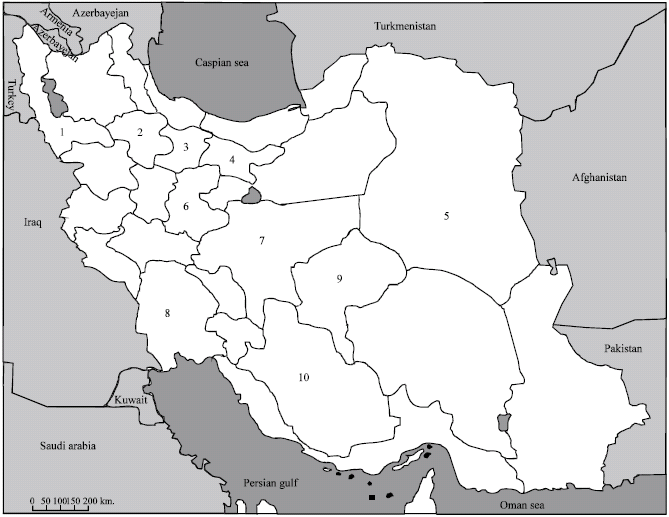
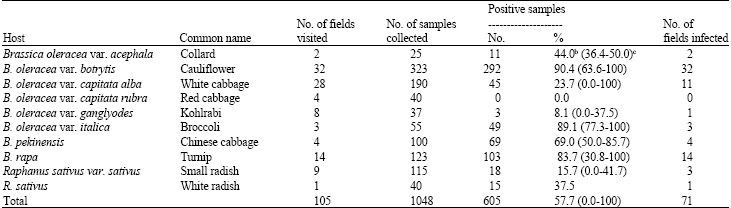
Occurrence and Distribution of Cauliflower mosaic virus on Cruciferous Plants in Iran
Department of Plant Protection, College of Agriculture, Tehran University, Karaj, Iran
Ali Ahoonmanesh
Department of Plant Pathology, College of Agriculture, Esfahan University of Technology, Esfahan, Iran
Gholam Hossein Mosahebi
Department of Plant Protection, College of Agriculture, Tehran University, Karaj, Iran
Reza Pourrahim
Department of Plant Virology, Plant Pests and Diseases Research Institute, P.O. Box 19395-1454, Tehran, Iran
Ali Reza Golnaraghi
Department of Plant Protection, Islamic Azad University, P. 0. Box 14515-775, Tehran, Iran