Research Article
Unearthed New Indel in the Waxy Gene in Glutinous Rice Landraces from Thailand
Department of Biotechnology, Faculty of Technology, Mahasarakham University, Maha Sarakham Province, 44150, Thailand
LiveDNA: 66.35236
Thailand is in mainland Southeast Asia and is part of an area referred to as the glutinous rice zone1. In ancient times (B.E. 1353-1707), northeastern Thailand and Laos were in the Lan Xang Kingdom in Southeast Asia, meanwhile (B.E. 1896-2250), northern Thailand was in the Lan Na Kingdom2. The Lan Xang and Lan Na Kingdoms had ethnic groups categorized as hill tribe peoples and lowland people. The hill-tribes people, Hmong, Mien, Khamu and Karen, lived in northern Thailand and Laos and were ethnically dominant in Laos and northern Thailand. These areas also included: Tai Dam (Black Tai), Tai Daeng, Tai Lu, Tai Yuan and Tai Phuan3,4. These Tai’ethnic groups have been characterized by their consumption of glutinous rice as a staple food and their rich culture that has played a key role in ritual and ceremonial festivities5. Furthermore, in the northeastern region of Thailand and Laos, archaeological evidence indicates that around 1,800-2,500 years ago rice cultivation at Non-Nok Tha and The Old City Wall in Vientiane’s archaeological sites exhibited imprints of round-type grain hull in brick which were more likely glutinous rice6,7. In Thailand, there are more than 3,000 traditional glutinous rice cultivars and were also developed by collection and selection from glutinous rice landraces, such as Muey Nawng 62 M, Daw Nahng Nuan91, Niaw San Pa Tawng and Khao Pong Krai. At present, in northern and northeastern Thailand, most glutinous rice landraces have disappeared from local farms and have been replaced by improved glutinous rice such as RD 6, Sakon Nakhon, RD 20, Khiaw Ngooo8974. An Example of rice grain morphology of glutinous rice landrace is shown in Fig. S1 (additional data).
Starch in rice grains is composed of amylose and amylopectin. Amylose is synthesized by the Granule-Bound Starch Synthase I (GBSS I) enzyme. This enzyme is encoded by the Waxy (Wx) gene8. Glutinous rice grain was characterized by a lack of amylose in endosperm tissue9. Based on the germplasm accession data of glutinous rice conserved in the International Rice Genebank, Sattaka et al.10 reported that glutinous rice was grown in 37 countries worldwide. In China, an ethnic group called Dong grows Kam fragrant glutinous rice and shares the consumption with people in Southeast Asian countries11.
DNA markers have been used extensively to characterize germplasm, evaluate genetic diversit and provide important information in population genetics and evolution. Within Asian glutinous rice, several studies have reported Single nucleotide polymorphisms (SNPs) in Waxy gene1,12-16.
The promoter region and 5’ untranslated region (5’UTR) have played an important role in gene expression regulation in rice such as OsMT2b gene in rice embryo17, alcohol dehydrogenase gene18. Besides, SNPs in the 5’UTR of rice Waxy gene from Bangladesh shed light on the evolutionary pattern line in the general patterns of South and Southeast Asia15. Based on the significance of glutinous rice that plays an important role in these areas, traditional practices are recorded history from archaeological sites.
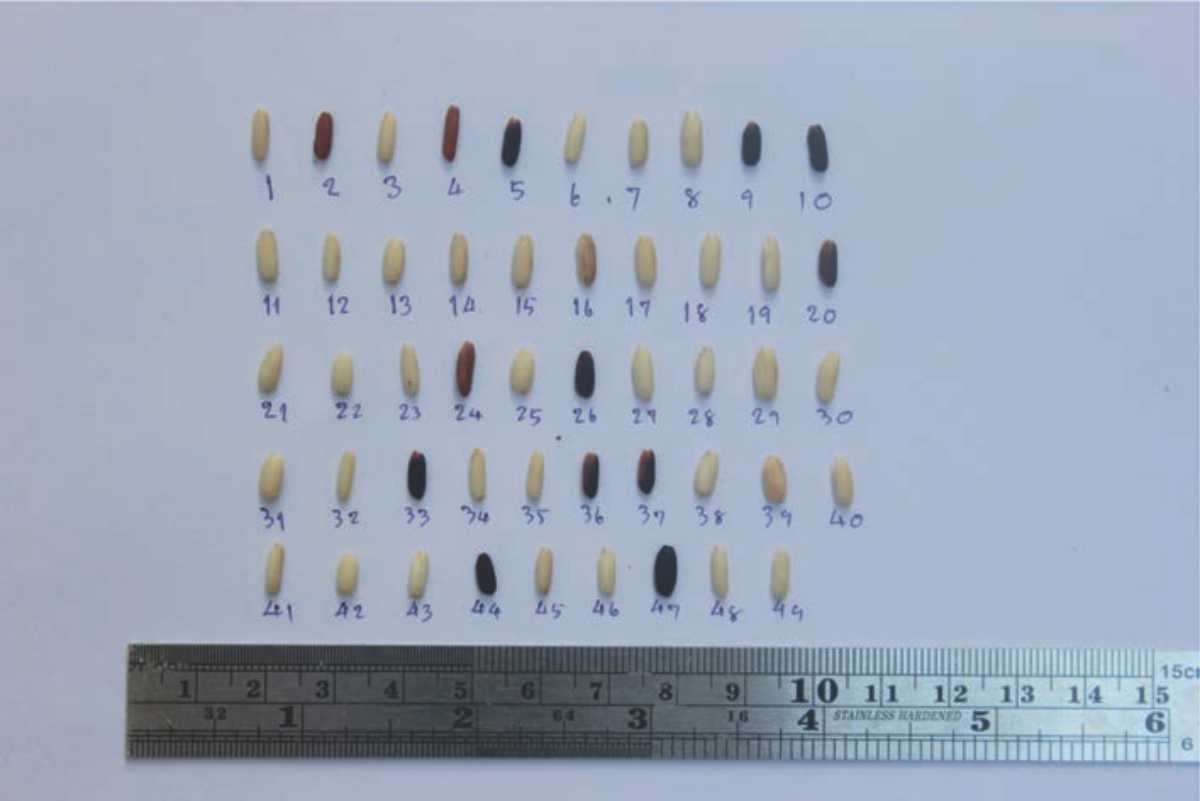 |
| Fig. S1: | Grains of 49 accessions of glutinous rice landraces collected from northeastern region Thailand, some of them used in this study |
It is a molecular characterization of these tropical glutinous rice cultivars that should be investigated further to help generate knowledge by applying molecular marker technology. Based on the waxy gene, it is noteworthy that there are some accessions of glutinous rice germplasm in Thailand that have been reported in previous report19,20. However, glutinous rice has not been given any serious study time resulting in only a few reports on genetic variation of glutinous rice. Besides, the study of Olsen and Purugganan1 used a small number of glutinous rice samples from Thailand and Laos (6 accessions).
Therefore, the main purpose of this study is to further characterize nucleotide variation in the promoter region and 5’UTR of the waxy gene (i.e., exon 1 and intron 1) in glutinous rice landraces and to determine the relationship between CT repeat alleles and a single-base substitution at a donor site of the leader intron in the waxy gene in a collection of present-day glutinous rice landraces from upland fields and paddy fields from Thailand and Lao PDR. As well as comparison with known haplotypes, which were deposited in the GenBank with aims to offer an alternative insight into their evolutionary pattern and to enhance germplasm use, which is the ultimate purpose of conserving the diversity of glutinous rice genetic resources.
Study area: This study was carried out at Walai Rukhavej Botanical Research Institute (WRBRI), Mahasarakham University, Thailand from October, 2018-September, 2019.
Rice materials: Seeds of fifty glutinous rice landraces both lowland paddy-rice and upland-rice cultivars were collected from ethnic groups (Khamu, Mien, Mhong) in northern Thailand (15 accessions) and Laos (1 accession) and lowland rice from farmers of northeastern Thailand (31 accessions) and Laos (3 accessions). The collections took place from 2004-2008. The samples were deposited in a laboratory as previously reported21. From 2017-2018, several surveys were made to collect glutinous rice landraces or accessions. Names, sources and localities and characteristics of the samples are listed in Table 1. Rice cultivar’s name corresponded to the Rice Department, Ministry of Agriculture and Cooperation, Thailand.
All seeds of glutinous rice samples were assessed at a low level of amylose by using an I2-KI assay (2.5 g KI, 250 mg I2, 125 mL distilled water). Waxy grain phenotype can be readily distinguished by iodine staining and waxy phenotypes usually stain reddish brown22.
For DNA extraction, five seeds of each rice accession were germinated and grown in pots at a greenhouse at Mahasarakham University, Thailand. Fresh leaves from a 1-month-old seedling were harvested for DNA extraction using the CTAB method as described by Doyle and Doyle23.
PCR and DNA sequencing: Polymerase Chain Reaction (PCR) amplification was performed by using two sets of primers as previously used in experiments by Olsen and Purugganan1.
The two primers set were as follows: W×U1F (5’-GCCGA GGGACCTAATCTGC-3’), W×U1R (5’-TGGTGTGGGTGGCTATT TGTAG-3’) and W×2Fa (5’-GCCCCGCATGTCATCGTC-3’), W×2R (5’-GTTGTCTAGCTGTTGC TGTGGA-3’).
The PCR was carried out using 0.5 units of Taq DNA polymerase, 0.5 μg of genomic DNA, 10 pmol of each primer, 1.5 mM MgCl2, 0.1 mM dNTPs in a final volume of 40 μL. The PCR conditions were 94°C (2 min); then 30 cycles of 94°C (30 sec), 55°C (30 sec), 72°C (2 min) and a final extension of 72°C (5 min). PCR products were analyzed by 1% agarose gel electrophoresis and visualized by Electrophoresis Gel Photo Documentation System (Bio-Rad 2000, Germany). The PCR products were further purified using Wizard® PCR Preps DNA Purification System (Promega, Madison, USA).
PCR products were directly sequenced using the primer W×U1F (5’-GCCGAGGGACCTAATCTGC-3’) for the promoter and 5’UTR and W×2Fa (5’-GCCCCGCATGTCATCGTC-3’) for determining G/T SNP in the junction exon 1-intron 1. PCR products were sequenced by using Applied Biosystems of DNA Sequencing Services of 1st BASE, Science Park Road, The Gemini, Singapore Science Park II, Singapore. DNA sequences from this study were deposited in GenBank (accession No. MK508875.1-MK508898.1).
DNA sequence analysis: Sequencing chromatograms of DNA sequences of all examined rice accessions were proofread to ensure that all ambiguous sites were correctly called and to determine the overall quality of data. The promoter region and the 5’UTR DNA sequences of a waxy gene of rice samples were aligned using Clustal W (http://www.genome.jp/tools/ clustalw/)24 by comparing with a previously reported haplotype, haplotype G (GenBank: AY136773.1) which is found in a sample of glutinous rice and were representative Indica and tropical Japonica (Javanica) from Thailand and Laos1. SNPs and base substitution (insertion and deletion) were identified by visual inspection of the multiple DNA sequence alignments.
| Table 1: List of glutinous rice landraces, improved glutinous rice cultivars and non-glutinous rice used in this study and their DNA sequence information | |||||||
| Cultivar’s name | Ethnic group/Field condition | Locality/province* | INDEL of (promoter region) | (CT)n | (T)n | T/A SNP position 2412 | G/T SNP at splice site intron 1 |
| Hao Mafai | Thai/upland | NE/Mukdahan | GG-ACC- | 17 | 10 | GT(A)7 | T |
| Hao Kaen Du | Thai/upland | NE/Mukdahan | GG-ACC- | 17 | 10 | GT(A)7 | T |
| U Kham | Thai/lowland | NE/Mukdahan | GG-ACC- | 18 | 10 | G(A)7 | T |
| Plah Kheng | Thai/lowland | NE/Mukdahan | GG-ACC- | 17 | 10 | G(A)7 | T |
| Perd Nam | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 18 | 10 | G(A)7 | T |
| Khao Yai | Thai/lowland | NE/Kalasin | GG-ACC- | 17 | 10 | G(A)7 | T |
| Leung Bun Mah | Thai/lowland | NE/Yasothon | GG-ACC- | 17 | 10 | G(A)7 | T |
| Pawng Aew | Thai/lowland | NE/Maha Sarakham | GG-ACC- | 17 | 10 | G(A)7 | T |
| E Pon | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 17 | 10 | G(A)7 | T |
| Khi Tom Hahng Nahk | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 17 | 10 | G(A)7 | T |
| Ma Yom | Thai/lowland | NE/Kalasin | GG-ACC- | 18 | 10 | G(A)7 | T |
| Man Ped | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 17 | 10 | G(A)7 | T |
| Nam Man Wua | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 18 | 10 | G(A)7 | T |
| Nam Pa | Thai/lowland | NE/Kalasin | GG-ACC- | 18 | 10 | G(A)7 | T |
| Pan Pae | Thai/lowland | NE/Kalasin | GG-ACC- | 17 | 10 | G(A)7 | T |
| E Puay | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 17 | 10 | G(A)7 | T |
| E Tia | Thai/lowland | NE/Mukdahan | GG-ACC- | 17 | 10 | G(A)7 | T |
| E Lub | Thai/lowland | NE/Yasothon | GG-ACC- | 17 | 10 | G(A)8 | T |
| E Khao Yai | Thai/lowland | NE/Kalasin | GG-ACC- | 17 | 10 | G(A)7 | T |
| Khi Tom | Thai/lowland | NE/Mukdahan | GG-ACC- | 18 | 10 | GT(A)7 | T |
| Khi Tom Yai | Thai/lowland | NE/Mukdahan | GG-ACC- | 17 | 10 | GT(A)7 | T |
| Kam Nai | Thai/lowland | NE/Kalasin | GG-ACC- | 17 | 10 | GT(A)7 | T |
| Ka Saen | Thai/lowland | NE/Maha Sarakham | GG-ACC- | 17 | 10 | G(A)8 | T |
| In Paeng | Thai/lowland | NE/Mukdahan | GG-ACC- | 17 | 10 | G(A)7 | T |
| Daw Nahng Nuan | Thai/lowland | NE/Mukdahan | GG-ACC- | 18 | 10 | G(A)7 | T |
| Daw Wi | Thai/lowland | NE/Mukdahan | GG-ACC- | 18 | 10 | G(A)7 | T |
| Dawk Mai Lai | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 17 | 10 | G(A)7 | T |
| Daw Gaset | Thai/upland | NE/Mukdahan | GG-ACC- | 17 | 10 | G(A)7 | T |
| Gaw Diaw | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 17 | 10 | G(A)8 | T |
| Plah Sew | Thai/lowland | NE/Sakon Nakhon | GG-ACC- | 17 | 10 | G(A)7 | T |
| Kai Noi | Hmong/lowland | Laos/Xiangkhouang | GG-ACC- | 17 | 10 | GT(A)8 | T |
| Mae Hahng | Laos/lowland | Laos/Vientiane | GG-ACC- | 17 | 10 | G(A)7 | T |
| Pae Daeng | Laos/lowland | Laos/Vientiane | GG-ACC- | 17 | 10 | G(A)7 | T |
| E Tia Kon Dam | Laos/lowland | Laos/Vientiane | GG-ACC- | 18 | 10 | G(A)7 | T |
| Peun Neon Klog | Khamu/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)7 | T |
| Bael Plao Chong | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)8 | T |
| Bael Cha Plau | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)7 | T |
| Bael Jao Blao | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)7 | T |
| Bael Leu | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)10 | T |
| Bao Bud | Mien/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)7 | T |
| Bael Dao Derk | Hmong/upland | N/Tak | GGGACC- | 17 | 10 | G(A)8 | T |
| Bael Di Nong | Hmong/upland | N/Tak | GG-ACC- | 17 | 10 | G(A)8 | T |
| Bael Jah | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | GT(A)7 | T |
| Bael Lao Da | Hmong/upland | N/Tak | GG-ACC- | 17 | 10 | G(A)8 | T |
| Biaw Gu | Mien/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)8 | T |
| Ja Ngai | Khamu/upland | N/Chiang Rai | GG-ACC- | 16 | 10 | G(A)7 | T |
| Gala | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 11 | G(A)11 | T |
| Khao Pong Krai | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)8 | T |
| Ram Tahng | Khamu/upland | N/Chiang Rai | GG-ACC- | 16 | 10 | GT(A)7 | T |
| Yim | Hmong/upland | N/Chiang Rai | GG-ACC- | 17 | 10 | G(A)8 | T |
| KDML105 | Thai/lowland | NE/Roi Et | GG-ACC- | 17 | 9 | G(A)7 | T |
| (non-glutinous, improved cultivar) | |||||||
| RD6 (improved cultivar) | Thai/lowland | NE/Mukdahan | GG-ACC- | 11 | 10 | GT(A)7 | T |
*N: Northern, NE: Northeastern, di-microsatellite (CT)n, mono-microsatellite (T)n, Single nucleotide polymorphisms (SNP) | |||||||
DNA sequences obtained from using the primer W×2Fa (5’-GCCCCGCATGTCATCGTC-3’) to determine G/T SNP at the waxy gene leader intron and the microsatellite CT repeats in the exon 1 were also observed by visual inspection. Neighbor-Joining (NJ) analyses25 was used to determine genealogical relationships among glutinous rice landraces which were implemented in MEGA 4 software26.
Nucleotide variation at the promoter region and 5’UTR of waxy gene: The samples of 50 glutinous rice landraces were examined for genetic variation in a promoter region, exon 1 and intron 1 that is a portion of the waxy gene. These cultivars were representative of the traditional glutinous rice diversity growing in these areas and passed from generation to generation.
Glutinous rice in Thailand and Lao PDR carried haplotype G1. DNA sequences of all glutinous rice samples were compared to the promoter region and the 5’UTR DNA sequence of haplotype G. Overall, two deletion G at position 11 (GG-) and C at position 15 (CC-) were observed in the promoter region of the 50 glutinous rice samples used in this experiment. The T/A SNP was observed in some glutinous rice landraces (Fig. 1). The nucleotide variation of these bases has not yet been reported and therefore it might be unique to glutinous rice samples from both tropical japonica and indica lowland rice of Thailand and Laos. Previously, new SNPs in Bangladeshi rice cultivars according to haplotype G were also observed by Shahid et al.15. This finding probably shed light on the genetic background of tropical japonica glutinous rice grown by ethnic groups in Thailand and Laos, which share their ancestry among glutinous rice landraces in these areas.
Variation of CT repeat alleles and mono-microsatellite T in glutinous rice landraces: The CT repeat alleles located in exon1 of the rice waxy gene, overall two classes, (CT)17 and (CT)18 were observed in the glutinous rice sample used in this study. This additional result supports the previous report that is most of the glutinous rice germplasm in Thailand and Lao PDR carried the two classes of CT repeat (i.e., (CT)17 and (CT)18) in both tropical japonica and indica types19,27. It might be concluded that 17 and 18 CT repeat alleles in glutinous rice germplasm in Thailand and Laos directly associate with glutinous (0-5% amylose). Whereas non-glutinous cultivar KDML105 (12-17% amylose) with 17 CT repeat allele had a low amylose content class (6-18% amylose). The 17 and 18 CT repeat alleles were also found in the US and European rice germplasm with low amylose classes13. Meanwhile, Bangladeshi rice had 17 CT repeat alleles in intermediate, low amylose content and waxy15. The Mono-microsatellite repeat T, (T)n at position 65-74 of the 5’UTR DNA sequences showed polymorphism. Almost of glutinous rice landraces had (T)10, while glutinous rice landrace ‘Gala’ from Hmong ethnic group exhibited (T)11.
G/T-SNP at the donor site of the first intron in glutinous rice: The fifty glutinous rice accessions samples were characterized for an SNP of the nucleotide at the first intron of the waxy gene. The result revealed that all glutinous rice examined had nucleotide T-SNP at the 5’ splice site in intron 1 of the waxy gene and could explain the glutinous phenotype or waxy. This finding supports the previous work in Thai glutinous rice by Prathepha19,20, glutinous rice from northern, Laos27. However, 11 Bangladeshi glutinous rice cultivars (Beruin) showed polymorphism in G/T SNP in splice donor site15. Furthermore, T/A-SNP was observed in intron 1 at position 2412. All glutinous rice landraces showed A SNP at the position, except a cultivar ‘Ram Thang’ from northern Thailand was identical to the reference DNA sequence (T-SNP).
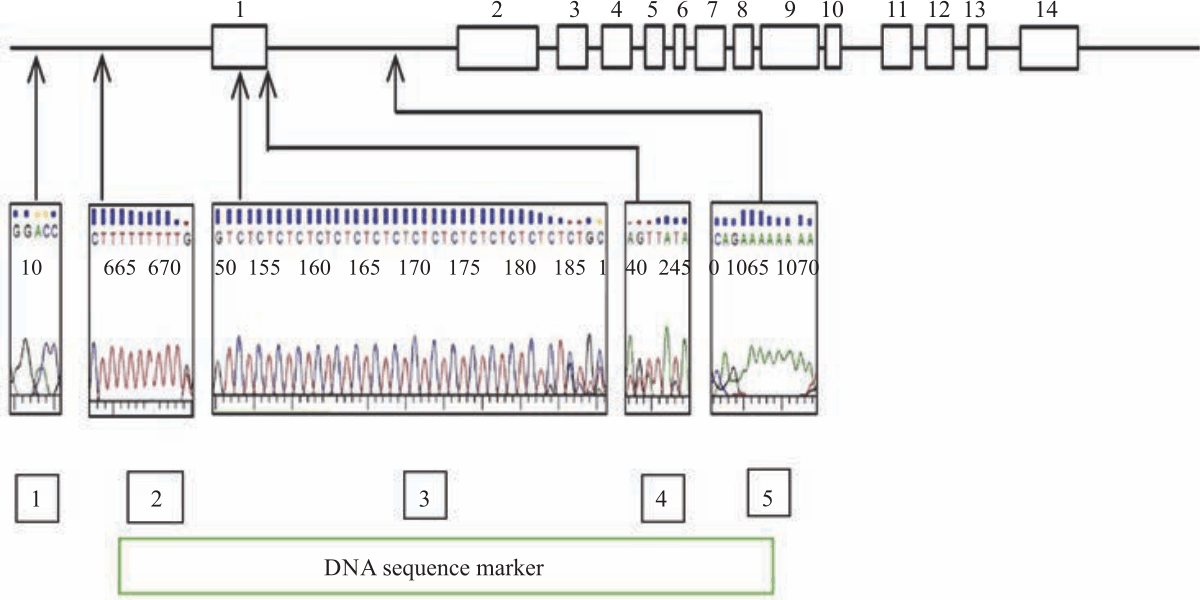 |
| Fig. 1: | Diagram of the Waxy gene, boxes and thick lines present exons and introns, respectively Chromatograms show the InDel (GG-ACC-)[1], mono-microsatellite (T)n at the 5’UTR [2], di-microsatellite (CT)n located in exon 1 [3], G/T SNP at the donor splice site [4] and T/A SNP at position 2412 in the first intron [5] |
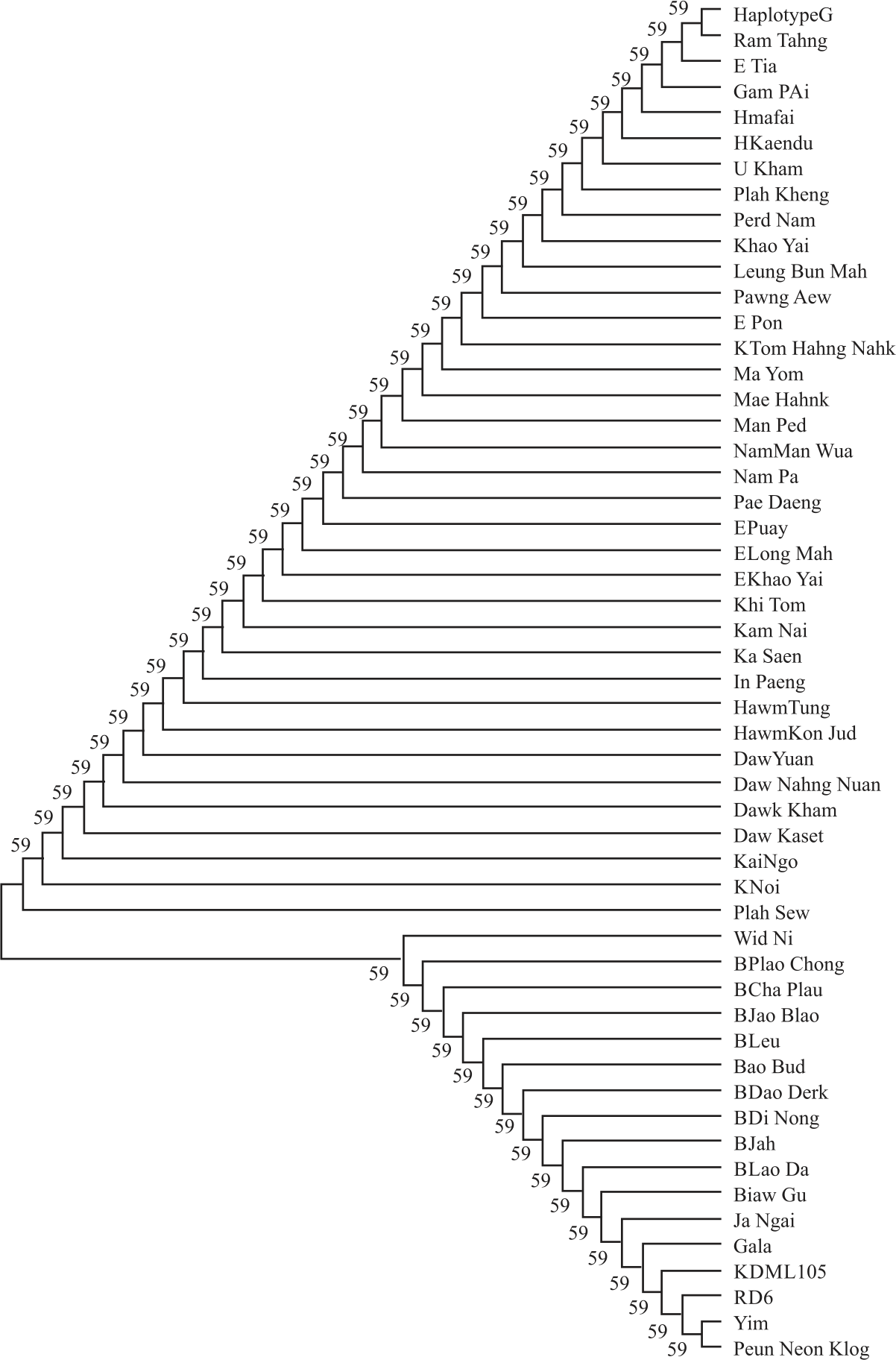 |
| Fig. 2: | Neighbour-joining Tree inferred from DNA sequences of 50 glutinous rice landraces Number in branches correspond to a bootstrap value |
Genetic structure of glutinous rice landraces: According to previous studies, glutinous rice exhibited a characteristic of 23 bp duplication in exon 2 of the waxy gene28. Besides, based on DNA markers, glutinous rice landraces grown in the upland field were categorized as tropical japonica type, whereas indica types were grown in lowland (paddy) field21,28. Previously, glutinous black rice in Thailand and Laos showed the 23 bp duplication and DNA markers were also applied to classify glutinous black rice into tropical japonica and indica types29.
A neighbour-joining tree constructed using the concatenated DNA sequences from all rice samples is shown in Fig. 2, which supports the expected pedigree of two types of rice (lowland indica versus upland tropical japonica), which are likely to have been classified using isozyme system30. Although the ancestry of rice landraces grown by ethnic groups is unclear, their position on the tree supports the notion that tropical japonica forms (upland rice in northern Thailand and Loa PDR) of japonica are more related to each other than to the indica forms (lowland rice from northeastern Thailand) and suggesting that these tropical japonica cultivars share a common heritage with japonica forms.
According to the promoter region and the 5’UTR DNA sequences of a waxy gene, the genetic variation observed in glutinous rice samples used in this study revealed only some InDel, reflecting intensive selection by local farmers. The genetic variance in these glutinous rice landraces can be utilized in a breeding program with crossbreeding. DNA markers found in these samples were CT repeat alleles and several T repeats, these markers could be used as the marker-assisted selection for improving grain trait with high cooking quality. Despite glutinous rice with gain starch contains low amylose content, each glutinous rice variety gives different eating quality characteristics to the processed rice products31. Besides, the importance of the genetic relationship among glutinous rice landraces in avoiding inbreeding within a breed can prevent excessive increases in the relationship in a breeding program.
This study illustrated DNA variation in the promoter region and the 5’UTR of a waxy gene, CT repeat alleles and G/T SNP at the 5’ splice site in intron 1 of a waxy gene. Rice samples used in this study were present time glutinous rice landraces of local farmers or ethnic groups in Thailand and some accessions from Lao PDR. New InDel and SNPs were found in the DNA sequence of glutinous rice landraces. Microsatellite CT repeat, (CT)17 and (CT)18 were two alleles found in samples. All glutinous rice landraces have T-SNP at the splice site in intron 1. The results of this study could shed light on genetic variation in glutinous rice landraces in this region. Furthermore, this additional data could be beneficial for making a plan for the conservation of these glutinous rice germplasm of ethnic groups and seeking a way to scaling up to develop new varieties for economy-based glutinous rice landrace.
This study discovers DNA variation in the Waxy gene that can be beneficial for germplasm evaluation and evolutionary studies for better strengthening the conservation and utilization of genetic diversity of glutinous rice landraces. This study will help the researcher to uncover the critical areas of genetic variation in glutinous rice in other areas that researchers were not able to explore.
This work was fully supported by Mahasarakham University with grant number {MSU-2563}. The author would like to thank V. Pilap for laboratory assistance and P. Dulfer for critical reading and improving English.