Research Article
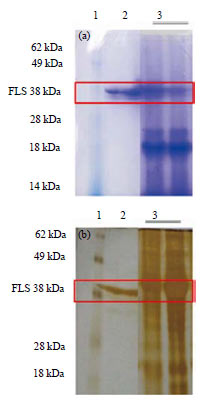
Determination of Quercetin and Flavonol Synthase in Boesenbergia rotunda Rhizome
Department of Pharmacology and Clinical Pharmacy, Faculty of Pharmacy, Padjadjaran University, Jalan Raya Bandung-Sumedang km 21, Jatinangor, 45363 West Java, Indonesia
Irma Melyani Puspitasari
Department of Pharmacology and Clinical Pharmacy, Faculty of Pharmacy, Padjadjaran University, Jalan Raya Bandung-Sumedang km 21, Jatinangor, 45363 West Java, Indonesia
Ronny Lesmana
Division of Molecular Physiology, Department of Anatomy, Physiology and Cell Biology, Faculty of Medicine, Padjadjaran University, Jalan Raya Bandung-Sumedang km 21, Jatinangor, 45363 West Java, Indonesia
Jutti Levita
Department of Pharmacology and Clinical Pharmacy, Faculty of Pharmacy, Padjadjaran University, Jalan Raya Bandung-Sumedang km 21, Jatinangor, 45363 West Java, Indonesia
LiveDNA: 62.1290