Research Article
Dynamic of Bacterial Diversity in Ileum Digesta Under Water Supplements of Antibiotics and Probiotics
Department of Animal Production, Mutah University, Karak Governorate, Jordan
LiveDNA: 962.4295
It is well-known that newly hatched birds lack digestive microflora and are highly susceptible to enteropathogen colonization and infection. In addition, digestive bacterial populations changes by many factors such as age and diet1,2. Several studies have suggested that the modulations of poultry feed or supplementation are responsible for the change of the intestinal microbiota composition.
Intestinal bacteria play an important role in the nutritional, physiological, immunological and also protective functions of the birds3. They modify the intestinal environment by reducing the pH, supplying digestion enzymes and increasing enzymes activity in the gastrointestinal tract of the birds3,4. The understanding of intestinal microbiota that comes from culture-based methods may be incomplete and inaccurate, because only 10-60% of the total intestinal tract bacteria are cultural5. The recent development of molecular technologies based on the 16S ribosomal RNA (rRNA) gene has resulted in detailed detections of microbial species in intestines6,7. The most common applied technology for sequencing rRNA gene of bacteria is bacterial tag-encoded 16 FLX ampliconpyrosequencing (bTEFAP). This technique is based on a partial ribosomal amplification followed by pyrosequencing8-10. This study advances the new knowledge of the microbiota diversity in organism health, disease and to develop viable therapeutic strategies. In the present study, we used bTEFAP16S ribosomal RNA (rRNA) gene based technology to investigate the diversity of microflora in the ileum.
Origin and collection of ileum samples: Thirty two samples were collected from the flock (four samples from each treatment) to estimated DNA of ileum microbial diversity. The birds were randomly chosen from Six hundred 1-day-old Ross broilers reared in Animal research station at King Saud University in summer of 2015. The treatments were classified into four treatments; tape water (control), tape water with antibiotics, water with probiotics and water with antibiotics and probiotics for 28 days of bird age. Primalac® is a multi-strain of probiotic in dry white powder form containing Lactobacillus Acidophilus Lactobacillus Casei, Bifidobacterium Thermophilum and Enteroccocus Faecium. While, The antibiotic which was used in our experiment is Pulmotil®/Tilmicosin®.
Each 28-days-old bird was killed and the ileum was removed. One gram of the ileum digesta was taken and suspended in 9 mL of PBS (Phosphate buffer solution) (pH 8.0) using stomacher blender (Seward Medical, London,UK) for 2 min. One mililiter of aliquot of each sample was removed into eppendorf tube and then frozen at -20°C until DNA extraction performed. The sampling and animal handling procedures followed the Implementing Regulations of the Law of Ethics of Research on Living Creatures (Saudi Arabia National Committee of Bio Ethics-http://bioethics.kacst.edu. sa/?lang= en-US) with the approval of the King Saud University Animal Ethics Committee.
DNA extraction: The total genomic DNA was extracted from 1 g digesta of ileum using a QIAamp stool DNA mini kit as recommended by the manufacturers (Qiagen, Valencia, CA). The extracted DNA samples were quantified using a The SPECTROstar® Nano-spectrophotometer from BMG company (BMG Labtech Ltd, UK) Nanodrop (Varian Techtron AA6) for measuring concentration and purity at absorbance ratio of 260/20811.
Bacterial tag-encoded 16 FLX ampliconpyro sequencing (bTEFAP): The bTEFAP of 16S rRNA sequencing services were provided by Genewiz Company, United States for bacterial identification. Specific bacterial specific primers suitable for microbial identification and metagenomic studies were included in PCR reactions12. All DNA samples were then subjected to PCR reactions (Varian Techtron AA6) with those 16S universal Eubacterial primers. All related and follow up procedures were performed following 454 Roche Genome Sequencer FLX System as detailed by Genewiz company. The resulted sequencing of each sample was processed using different custom scripted bio-informatics pipeline. The major bio-informatics pipeline used for to identify sequenced genome of bacterial isolates was BLAST database. The sequence of each sample was separately lunched into BLAST database for finding the closest match with best bacterial identification level at 98%. This identification level was defined "Threshold value" at and above which an agreement for species identification can be obtained.
Statistical analysis: Data were statistically analyzed by three-way ANOVA using the General Liner Models procedure of SAS, ver 9.213. The following model was used in order to investigate the tretaments effects in the experiment a complete random design14:
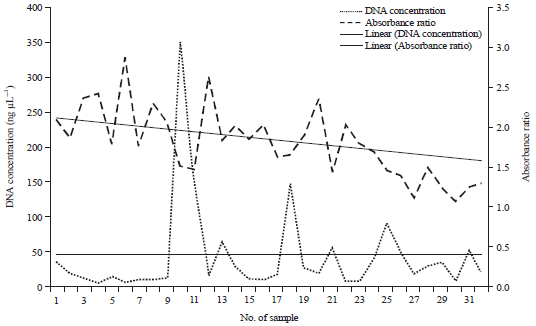 | |
| Fig. 1: | DNA quantification of all samples using Nano-DNA spectrophotometer |
![]()
where, Yijk is the individual observation, μ is the experimental mean, Ti is the effect of jth water supplement, eijk is random error.
DNA quantity and quality: The DNA concentration of the samples was determined for optimal sequencing and prior to the PCR amplification. The results of DNA concentration showed wide range 10-350 ng μL–1 (Fig. 1). This difference in the concentration was mainly due to handling and procedure of DNA extraction. On the other hand, quality of extracted DNA was also evaluated using the ratio at A260/A280 (Fig. 1). As a general rule, the quality of DNA is established by the ratio of absorbance at 260 nm to the absorbance of 280 nm. All samples absorbance ratios were averaged 1.8 and ranged from 1.0-2.6 as shown in Fig. 1. Cleaning DNA was then performed for those samples with outlier ratios.
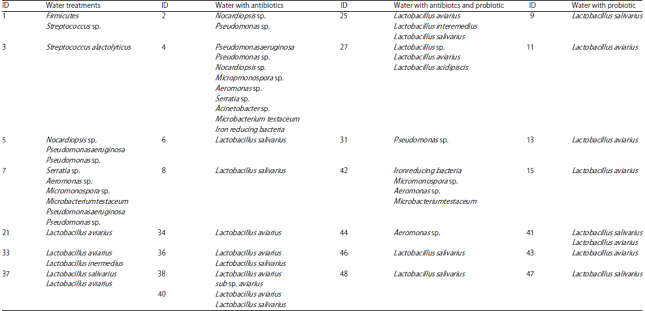
Ileum bactierial identification and diversity: The results of Table 1 showed the indentified bacteia populations as a response to treatments. These results were a summary of BLAST results for each sample. For example, Sample ID 1, 2, 21 and 9 were Water, Water with Antibiotics, Water with Probiotic and Water with Antibiotcs and Probiotic, respectively. They clearly showed a different bacterial population (Table 1). For first treatment, Sample ID 1, 3, 5 and 7 contained Firmicutes and pathogenic bacteria (Streptococcus sp., Streptococcus alactolyticu, Nocardiopsis sp., Pseudomonas aeruginosa, Pseudomonas sp., Serratia sp., Aeromonas sp., Micromonospora sp., Microbacterium testaceum, Pseudomonas aeruginosa, Pseudomonas sp.) whereas, Sample ID 21, 33, 37 contained non-pathogenic bacteria (Lactobacillus aviarius, Lactobacillus inermedius, Lactobacillus salivarius). In second treatment, Water with Antibiotics, Sample ID 2, 4, showed a content of pathogenic bacteria (Nocardiopsis sp., Pseudomonas sp., Pseudomonas aeruginosa, Pseudomonas sp., Nocardiopsis sp., Micropmonospora sp., Aeromonas sp., Serratia sp., Acinetobacter sp., Microbacterium, testaceum Iron reducing bacteria) whereas, Sample ID 6, 8, 34, 36, 38 and 40 contained only useful species of Lactobacillus salivarius and aviarius. In third treatment, Sample ID 25, 27, 46 and 48 had the following different Lactobacillus sp.; Lactobacillus aviarius, Lactobacillus interemedius, Lactobacillus salivarius and Lactobacillus acidipiscis. On the other hand, in same treatment, Sample ID 31, 42 and 44 had the following pathogenic species; Pseudomonas sp., Iron reducing bacteria, Micromonospora sp., Aeromonas sp. and Microbacteriumtestaceu. Finally, the last treatment -Water with Antibiotcs and Probiotic- showed useful nonpathogenic bacteria for all samples. They were Lactobacillus salivarius and Lactobacillus aviarius. It is good to note that this later bacterial species, Lactobacillus aviarius, is very useful bacteria species which have enhancing characteristics on immunity system of the organism.
On the other hand, the identified bacterial species presented in Table 1 were earlier classified as Operational taxonomic units (OTUs) defined as clones that shared 98% or greater sequence similarity. Afterwards, dendrogram construction for each sample was performed representing the most prevalent OTUs from each sample.
| Table 1: | Name of identified bacteria in the four treatments samples |
 | |
The dendrogram constructed showed the identified bacteria species in phylogenetic tree. As an example of phylogentic tree analysis utilized in identification bacterial species diversity, the taxonomy of identified bacterial species populations in Sample ID 7 of water treatment is shown in Fig. 2. The other phylogenetic trees of the remaining selected samples are presented in Fig. (3-5).
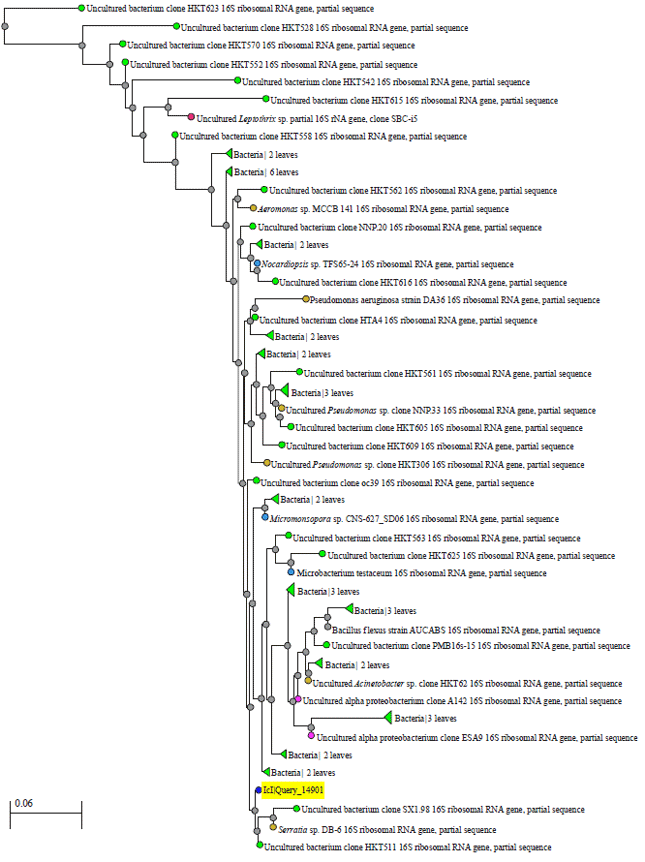 | |
| Fig. 2: | Phylogentic tree of identified bacterial species in sample ID 7 of water treatment |
The phylogenetics classified DNA sequencing according to their similarities and difference. It can be noted from the figures that the identified bacterial species a tree-like pattern that described the evolutionary relationships and how treatments were diverse bacteria species.
 | |
| Fig. 3: | Phylogentic tree of identified bacterial species in sample ID 7 of water and antibiotic treatment |
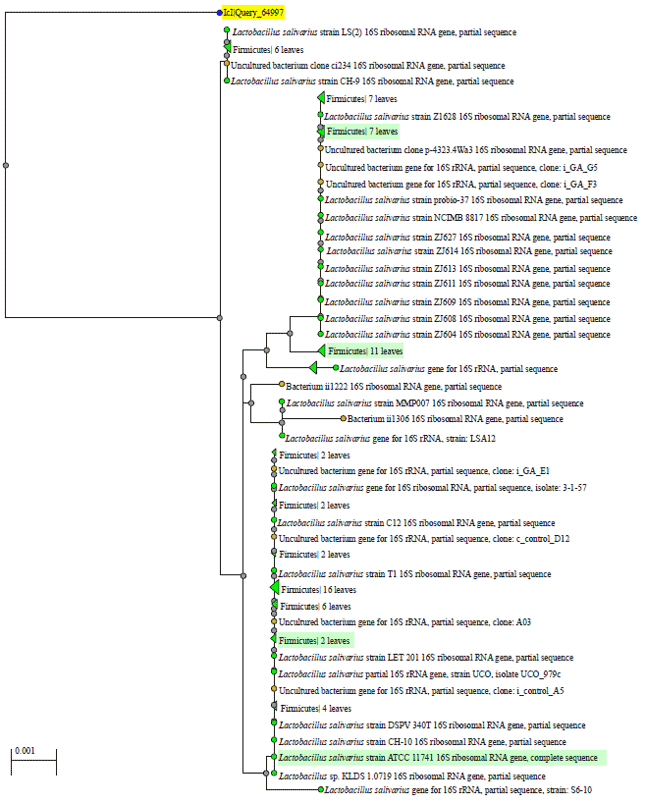 | |
| Fig. 4: | Phylogentic tree of identified bacterial species in sample ID 47 of water and probiotic treatment |
In the present study, the bacterial species in broiler ileum were identified based on their isolated DNA using bTEFAP procedures. First the DNA quantification and qualification was determined for optimal sequencing and prior to the PCR amplification. The wide range of DNA concentration (10-350 ng μL–1, Fig. 1) was due mainly to handling procedure of DNA extraction. In general, the best performance with PCR amplification of using commercially available kits occurred within a fairly narrow of input. Typically DNA amount of 5-2.5 ng μL–1 was enough to perform PCR reactions11,15.
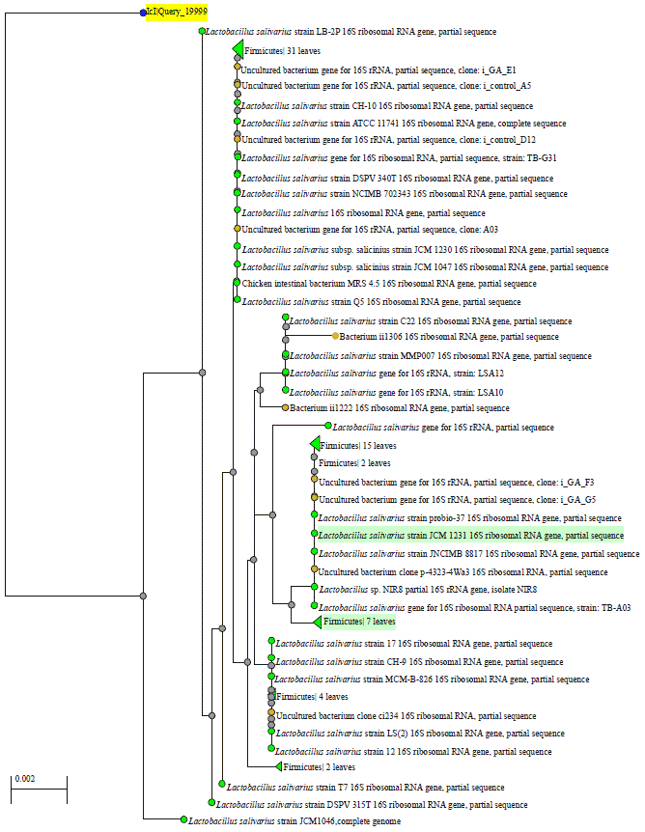 | |
| Fig. 5: | Phylogentic tree of identified bacterial species in sample ID 48 of water , antibiotic and probiotic treatment |
In addition as a general rule, the quality of DNA is established by the ratio of 260:280 nm absorbance of 280 for highly purified DNA, was averaged 1.8 which in accordance with the recommended ratio11.
The Ileum bactierial identification discovered which species existed as a result of giving antibiotics and probiotics treatments in the diet.The notable results were observed for Water with Probiotic treatment in which Lactobacillus salivarius and Lactobacillus aviaries were dominant. However, Pathogenic bacteria species were mainly observed water treatments. This result was expected for such control based treatment where pathogenic bacterial population find suitable niche to grow and reproduce as long growth-promoting antibiotics and antiparasitic agents not available. The results of the other two treatments are somehow in need for further investigation and justification. For example, the antibiotics make shift in the bacteria but becoming increasingly resistant to antimicrobial agents, Probiotics, on the other side, exhibited useful nonpathogenic bacteria for all samples. They were Lactobacillus salivarius and Lactobacillus aviarius. It is good to mention that this later bacterial species Lactobacillus aviarius, was very useful bacteria specie which have enhancing characteristics on immunity system of the those birds. The similar results were very earlier reported basis on culture-based methods of enumerated anaerobic bacteria from the ilea of 14-day-old chicks. The predominant cultured flora of the ileum included Lactobacillus, Streptococcus, E. coli and eubacteria16,17. Another report generated by molecular detection, showed also similar findings with our results18,19. They reported that Lactobacillus and Streptococcus were accounting for 62.7 and 12.2% of the ileum micro flora, respectively, along with many species of Clostridiaceae and fairly low of proteobacteria. Our findings were similar, with Lactobacillus species solely accounting for Probiotc treatment and Streptococcus and Pseudomonas accounting majority for other treatments. The constructed dendrogram classified the species based on similarities and difference in DNA sequencing. Noting a tree-like pattern that described the evolutionary relationships the treatments were successful in evolutionary distinct the bacteria species. In similar results, bacterial taxonomy for ileal communities showed Taxa significantly associated with cladogram under neutral situations20-22.
It is concluded that, the given diet treatments were made a shift in bacterial population diversity in ileum part of studied Broiler. The findings can be beneficial for the nutritional, physiological, immunological and also protective functions of both animal and human. This study promotes for a new approach in which replacing the use of probiotics as a prophylaxis in gastrointestinal infection and diseases instead of antibiotics. Knowing that the antibiotics make shift in the microbiota populations where infection may result, whereas, probiotics prevent diseases.
This study discovers the change in intestinal diversity as a result of giving antibiotics and probiotics in the diet. The expected results can be beneficial for the nutritional, physiological, immunological and also protective functions of both animal and human. This study will help the researcher to take decision on stop using antibiotics and start using probiotics as only alternative. The antibiotics make shift in the microbiota where infection may result. Thus a new approach may be practiced for giving probiotics as a prophylaxis in gastrointestinal infection and diseases, since pathogenic microorganisms are becoming increasingly resistant to antimicrobial agents.
The authors would like to extend their sincere appreciation to their Deanship of Scientific research at King Saud University for its funding of this research group with grant No. RG-1435-064.