Research Article
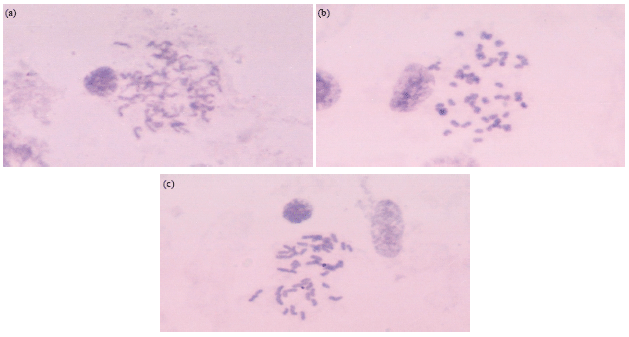
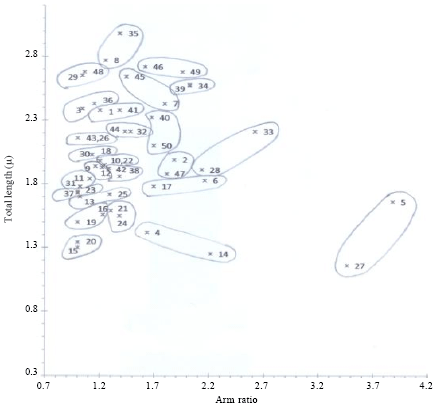
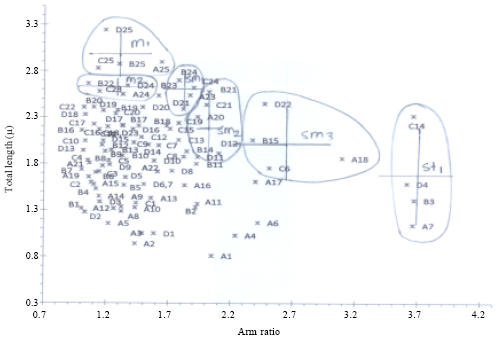
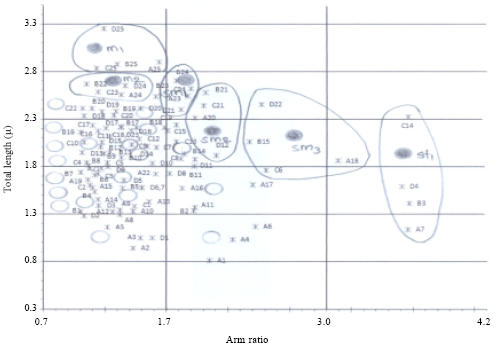
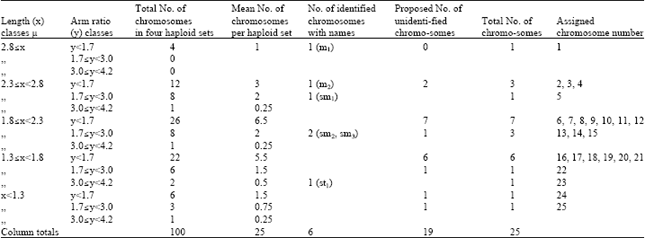
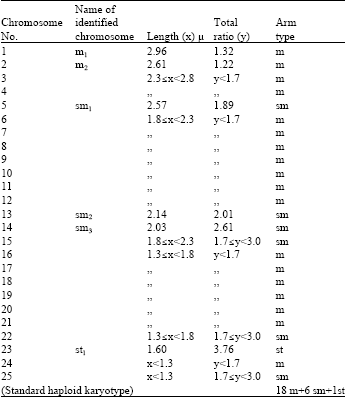
Chromosomal Studies and Quantitative Karyotypic Analysis of Rohu, Labeo rohita
Department of Fisheries Biology and Genetics, Bangladesh Agricultural University, Mymensingh-2202, Bangladesh
M. Belal Hossain
Biology Group, Faculty of Science, Universiti of Brunei Darussalam, BE1410, Brunei
M. Mokhlesur Rahman
Department of Fisheries Biology and Genetics, Bangladesh Agricultural University, Mymensingh-2202, Bangladesh
Md. Imran Hoshan
Department of Fisheries Biology and Genetics, Bangladesh Agricultural University, Mymensingh-2202, Bangladesh