Research Article
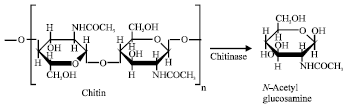
Novel Chitinase Genes from Metagenomic DNA Prepared from Marine Sediments in Southwest Japan
Department of Applied Bioresource Science, The United Graduate School of Agricultural Sciences, Ehime University, 3-5-7 Tarumi, Matsuyama, Ehime, 790-8566, Japan
S. Nagata
Department of Bioresources Science, Faculty of Agriculture, Kochi University, 200 Monobe, Nankoku, Kochi 783-8502, Japan
K. Ohnishi
Research Institute of Molecular Genetics, Kochi University, 200 Monobe, Nankoku, Kochi 783-8502, Japan