Research Article
Sequence Analysis of GDSL Lipase Gene Family in Arabidopsis thaliana
Department of Bioengineering, Zhengzhou University, 100 Science Road, Zhengzhou, Henan 450001, People`s Republic of China
INTRODUCTION
GDSL lipases widely exist in microbe and plant species. As an important gene family of lipases, GDSL lipases are active in hydrolysis and synthesis of lipids or esters. A serine-containing GDSL-motif close to the N-terminus, five conservative blocks and a Ser-Asp-His triad are included in the ammonic acid sequences (Upton and Buckley, 1995). GDSL lipase is generally made up of several ß-strands and a-helices arranged in alternate order and the substrate-binding pocket between the central ß-strand and long a-helix appears to be highly flexible. The flexible pocket brings conformational changes, so that the active sites exposure to the solvent and easily bind to substantive substrates. Possessing multiple functions, GDSL lipases are potentially applied to the food, flavor, fragrance, cosmetics, textile, pharmaceutical and detergent industry (Akoh et al., 2004; Ling et al., 2006).
Physiologically, plant GDSL lipases are generally considered to be mainly involved in the regulation of plant growth and development. Recently the research in this field becomes more and more attractive. Although several candidates from A. thaliana, Rauvolfia serpentina, Medicago, Hevea brasiliensis, Alopecurus myosuroides have been extracted, cloned and characterized (Brick et al., 1995; Oh et al., 2005; Ruppert et al., 2005; Pringle and Dickstein, 2004; Arif et al., 2004; Cummins and Edwards, 2004), such understanding of GDSL lipases is still limited. Comparatively, complete sequencing of the Arabidopsis genome accelerates cloning and characterization of lipases and information of Arabidopsis GDSL lipases deposited in the databases is rich and detailed. However, identification and functional analysis of them in databases or literatures is not enough or incomplete. Although a small number of Arabidopsis GDSL lipases (designated as AGLs) have been phylogenectically analyzed and aligned by Akoh et al. (2004), further comprehensive information is required by performing gene structure, multiple alignment, conserved motif or domain search and analysis of subcellular localization.
This study was conducted at Zhengzhou (China) in the year of 2006-2007.
The original set of the AGL cDNAs was searched at the Arabidopsis Information Resource [http://www.arabidopsis.org], then compiled and translated into amino acid sequences by Vector NTI Suite 8.0 [http://www.invitrogen.com]. The other protein sequences were retrieved from GenBank http://www.ncbi.nlm.nih.gov/Genbank/index.html] and EBI databases [http://www.ebi.ac.uk/interpro/].
All the gene structures were analyzed in website [http://www.ncbi.nlm. nih.gov/Entrez/query.fcgi?db=gene]. The motifs or domains of AGL proteins were analyzed using InterProScan [http://www.ebi.ac.uk/InterProScan/]. Sequences without GDSL-lipase domain or motif were removed. The left AGL protein sequences were aligned using Vector NTI Suite 8.0 to construct representative sequences. The gene loci distribution was performed by Chromosome Map Tool (http://www.arabidopsis.org/jsp/ChromosomeMap/tool.jsp). The phylogenetic analysis was carried out using the Neighbor-Joining method with MEGA3 http://www.megasoftware.net/]. Analysis of subcellular localization was performed for all the left AGL protein sequences by the target P program version 1.1 http://www.cbs.dtu.dk/services/TargetP].
Molecular analysis of multiple GDSL lipases and genes: Data mining was used to search all the cDNA sequences encoding AGLs. Using NCBI and TAIR database searches, 121 AGL cDNA sequences were obtained (data not shown). After sequence compilation, translation and alignment, thirteen cDNAs encoding other proteins were removed. Because two sequences (Acc. No. NM_118813 and NM_179120) are translated into one protein, the 108 cDNAs corresponding to 99 Atg loci putatively encode 107 GDSL lipase proteins.
As a result, two genes (Acc. No. NM_118813 and NM_179120) that encode one protein have different gene structures, so the structure and organization analysis of total 108 genes was performed in website. It indicated that the gene structures showed remarkable diversity. And it is likely that all of them except for one gene (Acc. No. AY058847) contain different numbers of introns. Of the analyzed genes, 73 genes consist of four introns; 11 and 12 have three and two introns, respectively. One gene (Acc. No. NM_101867) containing 13 introns is the most complicated and predicted to encode a high molecular weight protein of 1006 amino acid residues, while another one (Acc. No. NM_113550) with eight introns putatively encodes a much smaller protein of only 380 amino acids. And the other nine genes contain one or five introns. These introns and exons have the length of several decades of base pairs to over one kilo-base pairs.
Analyzed by Chromosome Map Tool, GDSL lipase genes are distributed in all chromosomes although not uniformly (Fig. 1). Forty seven genes exist in Chromosome 1 and 23 genes are distributed in Chromosome 5. Fourteen and Fifteen genes exist in Chromosome 2 and 3, respectively. Nevertheless, only eight genes are located in Chromosome 4. There are some regions with a high density of genes, such as the middle and bottom of Chromosome 1, the top and bottom of Chromosome 5. Furthermore, there are 12 cases of two or more genes arranged in tandem. For example, six genes (Acc. No. NM_202420, NM_106239-NM_106243) that encode extracellular lipases (EXL1-6, respectively) are arranged in tandem at the bottom of Chromosome 1, which is consistent with the previous study (Mayfield et al., 2001). In addition, some of them are likely duplicated in the areas of inter- or intra-chromosome, resulting in multiple gene copies in genome.
The obtained cDNA sequences were putatively translated into 107 amino acid sequences, consisting of 118-1006 amino acid residues, with the molecular weight from 12.9-109.1 kDa. Five blocks (block 1-5) containing the conserved sequences (PAIFVFGDSIVDTGNNN, TGRFSNGRLIXD, ALYLIXIGXNDY, LYXLGXARK XXVXGLXXXPLGCLP and YVFWXDXXHPTEXA, respectively) were determined by aligning the obtained sequences. Combining the previous reports (Oh et al., 2005; Akoh et al., 2004), we predict that the active serine, aspartate and histidine (in block 1 and 5, respectively) likely constitute the active triad.
Phylogenetic relationship was analyzed by the method of neighbor-joining. They were classified into three groups (Group 1, 2 and 3, Fig. 2), composed of 28, 40 and 39 members, respectively. Proteins designated as APG (Acc. No. AAL24235) and EXL1-6 were included in Group 2 and reported to be involved in flowering (Upton and Buckley, 1995; Mayfield et al., 2001) and GLIP1 (Acc. No. NP_198915, encoded by At5g40990) in Group 2 was involved in defense against the necrotrophic fungus Alternaria brassicicola as shown by Oh et al. (2005). Additionally, Arab-1 (Acc. No. NP_174188) belongs to Group 3.
Domain/motif structures of GDSL lipase proteins: Analysis by InterProScan indicated that all of the 107 proteins contained lipase-GDSL domain (PF00657). Whereas its location, number and kind are different, which leads to five classes of AGLs (class A-E) detailed in Table 1. It reveals that most AGLs in class A contain only lipase-GDSL domain close to N-terminus, which is consistent with characteristics of traditional GDSL lipases. Several to tens of amino acid residues in front of the lipase-GDSL domain likely shape into a signal peptide (SP). However, lipase-GDSL domain is located at the N-terminus and in the middle of two proteins (Acc. No. NP_196001 and AAL24235), respectively. Interestingly, three lipase-GDSL domains were found in NP_173441. Furthermore, one protein (Acc. No. NP_177718) contains two different domains (lipase-GDSL domain and 5`-nucleotidase/apyrase domain), suggesting that it might function as a GDSL lipase and a nucleotidase/apyrase. Although exist the above-mentioned differences, domain/motif structure of AGLs is comparatively uniform.
| Table 1: | The predicted domains located in AGL proteins |
 | |
| L: Lipase-GDSL domain, NA: 5`-nucleotidase/apyrase domain, AGL: Arabidopsis GDSL lipase | |
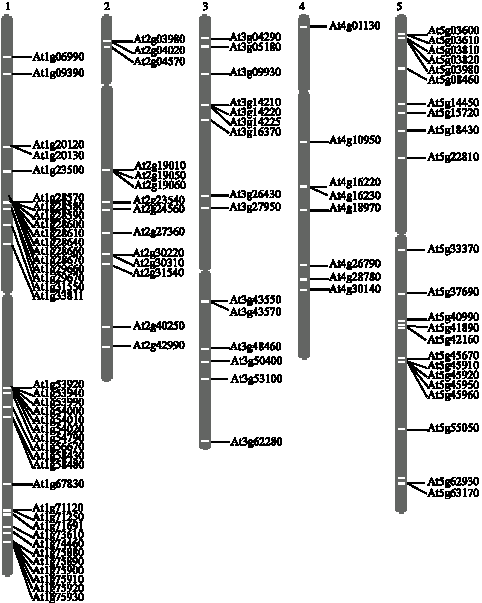 | |
| Fig. 1: | Loci distribution of Arabidopsis GDSL lipase genes in chromosomes. The loci distribution was performed in http://www.arabidopsis.org/jsp/ChromosomeMap/tool.jsp. |
Without enough resources of three-dimensional structure of plant GDSL lipases in the databases, presently three-dimensional structure could not be performed by SWISS-MODEL server.
Functionally, these lipases are potentially involved in multiple physiological roles. First of all, some play an important role in flowering. The anther-specific proline-rich protein (Acc. No. AAL24235) and extracellular lipases (EXL1-6) found in pollen coat have been identified (Brick et al., 1995; Mayfield et al., 2001). Secondly, some GDSL lipases function in disease resistance. For example, GLIP1 appears to trigger systematic resistance signaling
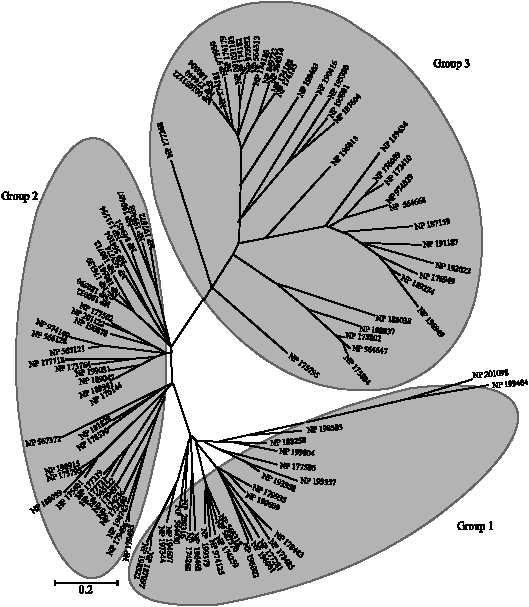 | |
| Fig. 2: | Phylogenetic analysis of Arabidopsis GDSL lipases the phylogenetic relationship of Arabidopsis GDSL lipase proteins were analyzed using the Neighbor-Joining method with MEGA3 |
in plant species when challenged by A. brassicicola (Oh et al., 2005). Additionally, other GDSL lipases potentially take roles in seed germination and other issues as well. This kind of protein in post-germinated sunflower (H. annuus L.) seeds shows fatty acid-ester hydrolase activity (Beisson et al., 1997). And an acetylajmalan esterase (designated as AAE, homologous to protein NP_174181) from Rauvolfi plays an essential role in the late stage of ajmaline biosynthesis (Ruppert et al., 2005). However, further evidences are required to support such predictions.
Prediction of subcellular localization: It reveals that 99 proteins in type 1 contain signal peptide, putatively involved in targeting to endoplasmic reticulum or subsequent transport through secretory pathway (Table 2). The signal peptides consist of 16 to 113 amino acid residues. A signal peptide was found in Arab-1 (Acc. No. NP_174188) and EXL1-6. Other four proteins (Acc. No. NP_199404, NP_201098, NP_199004 and NP_196001) were predicted to target to any parts in cells. Two proteins (Acc. No. NP_195980 and NP_173441) potentially target to mitochondria and chloroplast, respectively. While the subcellular localization of APG protein (Acc. No. AAL24235, also P40602) is still unclear. In Arabidopsis, a subfamily of genes encoding six extracellular lipases (EXL1-6) from pollen coat has been reported (Mayfield et al., 2001). Hereby, in addition to being extracellularly secreted, GDSL lipases with signal peptide probably target to different organelles or subcellular parts.
| Table 2: | Subcellular localization prediction of AGL proteins |
 | |
In conclusion, we analyzed the Arabidopsis GDSL lipase gene family of 108 members distributed in chromosome 1-5 and putatively encoding 107 GDSL lipase proteins. The gene structures, phylogenetic relationship, domain/motif organization and subcellular localization have been performed. It`s a superfamily of putative GDSL lipase genes potentially playing important roles in regulation of Arabidopsis growth and development, including seed germination, flowering and defense reactions. In the future, further data of precise subcellular localization, three-dimensional structure modeling, mutagenesis, over-expression/recombinant expression, substrate selectivity will be done to understand more about Arabidopsis GDSL lipases.