Research Article
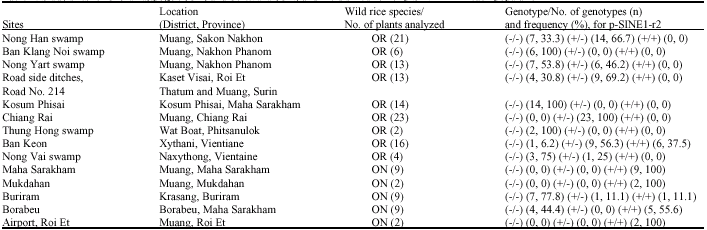
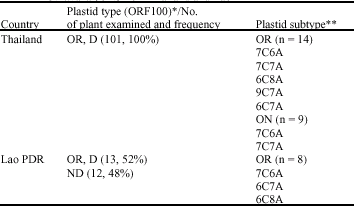
Genetic Variation of Wild Rice Populations from Thailand and the Lao PDR Based on Molecular Markers
Rice Genetics Research Unit, Walai Rukhavej Botanical Research Institute (WRBRI),
Mahasarakham University, Khamriang District, Maha Sarakham Province, 44150, Thailand