ABSTRACT
In order to determine stable bread wheat genotypes with high grain yield via a single parameter, field experiments were conducted with 20 genotypes for 4 consecutive years in two different conditions (irrigated and rainfed) in a completely randomized block design with three replications in each environment. Combined analysis of variance showed highly significant differences for the GE interaction indicating the possibility of selection for stable entries. The results of additive main effect and multiplicative interaction (AMMI) analysis revealed 10% of total variability was justified by the GE interaction which was 2.5 times more than that of genotypes. Ordination techniques displayed high differences for the interaction principal components (IPC1, IPC2 and IPC3), exhibiting that 83% of the GE sum of squares was justified by AMMI1, AMMI2 and AMMI3, i.e., 3.77 times more than that explained by the linear regression model displaying the relative efficiency of AMMI1 model in comparison with regression model. AMMI stability value (ASV) discriminated genotypes G6, G9, G14, G15, G17 and G18 as the stable entries. As stability per se is not a desirable selection criterion, because the most stable genotypes would not necessarily give the best yield performance, hence, simultaneous consideration of grain yield and ASV in a single non-parametric index entitled: Genotype Selection Index (GSI), discriminated G6 and G17 with general adaptability and high grain yield for rainfed and irrigated conditions which was in agreement with the results of biplot analysis.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/pjbs.2008.1791.1796
URL: https://scialert.net/abstract/?doi=pjbs.2008.1791.1796
INTRODUCTION
Bread wheat (Triticum aestivum L.) is a major food grain in Iran, therefore improving yield and yield stability is the primary objective of wheat breeding programs in this country (Ram et al., 2007).
GenotypexEnvironment (GE) interaction result in genotype rank changes from an environment to another, a difference in scale among environments, or a combination of these two situations. If relative performance of genotypes grown in different environments are different, then GE interaction becomes a major challenging factor to crop breeding programs (Mohammadi and Amri, 2008).
Adaptability is the result of genotypexenvironment interaction and generally falls into two classes: (1) the ability to perform at an acceptable level in a range of environment, general adaptability and (2) the ability to perform well only in desirable environments, specific adaptability (Farshadfar and Sutka, 2006; Solomon et al., 2008).
To identify wheat genotypes with wide or specific adaptation to different environments, multilocation yield trails are grown each year. These have led to empirical identification of superior cultivars, sum of which have been released in several counties (Asenjo et al., 2003; Basford et al., 2004).
The environments now involve a wide range of photoperiods and temperatures which could cause large genotypes (G) x Environment (E) interaction (GEI), especially in the semi-arid areas. Large real crossover-type GEI, especially among high yielding lines invalidates recommendations to farmers of the cultivar(s) giving the highest average yield across all test environments. Quantification of GEI and understanding its physiological bases are needed to breed efficiently for superior environments (Vergas et al., 2001; Thamson and Phillips, 2006). Most yield trails are used only to determine which cultivars give highest average seed yield, and therefore merit recommendation for planting by farmers. Multilocation yield trials facilitate quantification of the environment and GEI effects. However, a fact not generally recognized is that, in addition, every yield trial by analyzing processes that determine yield can inexpensively quantify the genetic, physiological and environmental controls that results in yield differences among cultivars, seasons and locations (Tarakanovas and Ruzgas, 2006). Various methods of GE interaction analysis exits, including parametric and non-parametric approaches.
The most widely used parametric method is the joint regression including regression coefficient (bi) and variance of deviation from regression (S2di) (Farshadfar and Sutka, 2006; Pourdad and Mohammadi, 2008).
Regardless of its advantage, joint regression analysis has the following disadvantages: (i) environmental index is not independent of the analyzed data, (ii) regression coefficients are biased because one assumption in the regression analysis is that independent variable (here, environmental mean) is measured without error, (iii) in this method it is assumed that a linear relationship exists between GE and environmental mean and (iv) relative stability of each pair of genotypes depends on the other genotypes in the experiment.
The ordinary form of ANOVA is an additive model and therefore describes only the main effect (Tabachnick and Fidel, 2007). Principal component analysis is a multiplicative model and has the opposite problem of not describing the additive main effects. Linear regression models combine additive and multiplicative components and thus analyze both main effects and interaction, but in general they confound the interaction with the main effects reducing its power for general significance testing (Farshadfar and Sutka, 2006).
The additive main effects and multiplicative interaction (AMMI) model is a powerful multivariate method to multienvironmental trails. This technique, also called FANOVA, incorporates both additive and multiplicative components into an integrated, powerful least square analysis (Farshadfar and Sutka, 2003; Mohammadi et al., 2007; Pourdad and Mohammadi, 2008).
AMMI is essentially effective where the assumption of linearity of responses of genotype to a change in environment is not fulfilled (Oliveira and Godoy, 2006), which is required in stability analysis techniques.
Complex relationships among location or among genotypes can be adequately in a scatter gram which show both the genotypes and environments simultaneously are called biplots (Mohammadi et al., 2007; Mahammadi and Haghparast, 2007).
The present research was carried out to quantify GE interaction effects on the yield and to determine stable entries within the genotype pool used in the study.
MATERIALS AND METHODS
In order to determine stable bread wheat genotypes field experiments were conducted for 4 consecutive years (1998-2001) in two different conditions (irrigated and rainfed) in College of Agricultural, Razi University, Kermanshah, Iran. Therefore 8 environments were created. The experimental layout at each environment was completely randomized block design with three replicates. The environments were considered as random factors while genotypes as fixed factors. Plots consisted of two 1 m rows spaced 20 cm apart. Average rainfall in the research station was 478 mm for each year. Maximum and minimum temperature was 44 and -27°C, respectively and the region was semi-arid. Combined analysis of variance, Bartlett, s test for additivity on grain yield and mean comparison with Duncan`s multiple range test were done using MSTST-C and SPSS statistical softwares. The additive main effect and multiplicative interaction (AMMI) analysis was performed using the model suggested by Crossa et al. (1991) as:
where, Yij is the yield of the ith genotype in the jth environment, μ is the grand mean, gi is the mean of the ith genotype minus the grand mean, ej is the mean of the jth environment minus the grand mean, λn is the square root of the eigen value of the Principal Component Analyses (PCA) axis, αni and γnj are the principal component scores for PCA axis n of the ith genotype and jth environment, respectively and Rij is the residual.
A biplot based on the Singular Value Decomposition (SVD) of GE contains only the GE interaction and can be referred to as a GE biplot. In contrast a biplot based on the SVD of G and GE contains only G plus GE and will be characterized as a GGE biplot (Weikai et al., 2000).
The GE biplot was projected for 20 genotypes tested at 8 environments. Clustering was computed for the genotype score using an agglomerate hierarchical algorithm based on Ward`s method (Farshadfar, 1998) and the result of cluster grouping for the genotype PCA score was projected in the biplot of PCA1 and PCA2 and the biplot of PCA1 and mean yield. The regression of yield for each variety on yield means for each environment was computed and parameters MS-REG, the contribution of each variety to the regression component of the treatmentxlocation (TL) interaction and MS-TL, the contribution of each variety to interaction MS, were estimated with the IRRISTAT program.
AMMI Stability Value (ASV): The ASV is the distance from the coordinate point to the origin in a two-dimensional of IPCA1 scores against IPCA2 scores in the AMMI model (Purchase et al., 2000). Because the IPCA1 score contributes more to the GE interaction sum of square, a weighted value is needed. This weight is calculated for each genotype and each environment according to the relative contribution of IPCA1 to IPCA2 to the interaction SS as follows:
where, ![]() is the weight given to the IPCA1-value by dividing the IPCA1 sum of squares by the IPCA2 sum of squares.
is the weight given to the IPCA1-value by dividing the IPCA1 sum of squares by the IPCA2 sum of squares.
The larger the IPCA score, either negative or positive, the more specifically adapted a genotype is to certain environments. Smaller IPCA score indicate a more stable genotype across environments.
Genotype Selection Index (GSI): Based on the rank of mean grain yield of genotypes (RYi) across environments and rank of AMMI stability value (RASVi) a selection index called GSI was calculated for each genotype which incorporate both mean grain yield and stability index in a single criteria (GSIi) as:
RESULTS AND DISCUSSION
The results of combined analysis of variance (Table 1) showed highly significant differences for genotypes, environments and GE interaction indicating the effect of environment in the GE interaction, genetic variability and possibility of selection for stable entries. As GE interaction was significant, therefore we can further proceed and calculate phenotypic stability (Farshadfar and Sutka, 2003; Farshadfar and Sutka, 2006). As Tukey`s test of additivity was not significant, it can be concluded that the effects are additive and additivity assumption of analysis of variance is provided (Snedecor and Cochran, 1987; Oritza et al., 2001). Bartlett test was significant exhibiting the heterogeneity of error variance, but as transformation of data caused missing some information and incorrect decision, hence no transformation exerted on the actual data (Hugh and Gauch, 1988).
Mean comparison using Duncan`s multiple range test revealed that maximum grain yield belonged to genotype number G20 (106.2 g) and minimum grain yield was attributed to genotype number G18 (61.43 g). The results of regression analysis (Table 2) exhibited that main effects of genotypes and GE interaction were relatively small and accounted for 4 and 10% of Total Sum of Square (TSS), respectively. Linear GE interaction was not significant and accounted for 22% of the variability in the GE interaction.
As a general rule the effectiveness of regression analysis is when 50% of the total sum of squares is accounted for by linear GE interaction (Hayward et al., 1993), hence regression analysis is not useful for stability analysis of genotypes (Wade et al., 1995). Nevertheless using regression analysis of Finlay and Wilkinson (1963) and Eberhart and Russell (1966), because of its wide application, it was concluded that genotypes number G4 and G17 have regression coefficient significantly greater than one with minimum deviation from regression indicating general adaptability for rainfed and irrigated conditions (Farshadfar, 1998; Farshadfar and Sutka, 1999).
| Table 1: | Combined analysis of variance for grain yield over different rainfed and irrigated conditions |
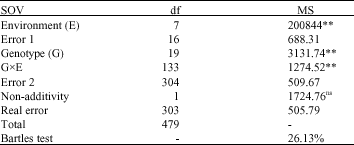 | |
| **Significant at 1% level of probability; ns: non-significant |
| Table 2: | Regression analysis of phenotype stability of bread wheat genotypes |
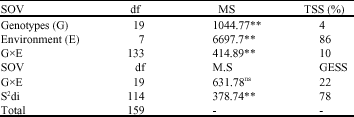 | |
| **Significant at 1% level of probability; ns: non-significant |
| Table 3: | AMMI analysis of grain yield in bread wheat genotypes over environments |
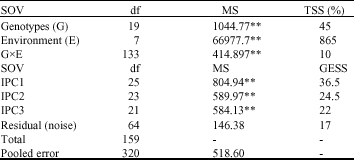 | |
| **Significant at 1% level of probability |
AMMI model and pattern analysis: In AMMI model, principal component analysis is based on the matrix of deviation from additivity or residual, while patter analysis employs both classification and ordination techniques. In this respect both the results of AMMI analysis, the genotype and environment will be grouped based on their similar responses (Gauch, 1992; Wade et al., 1995; Pourdad and Mohammadi, 2008).
Using ANOVA yield sum of square was partitioned into genotype, environment and GE interaction. GE interaction was further partitioned by principal component analysis (Table 3). The result of AMMI analysis indicated that 10% of total variability was justified by GE interaction, 86% by environments and 4% by genotypes.
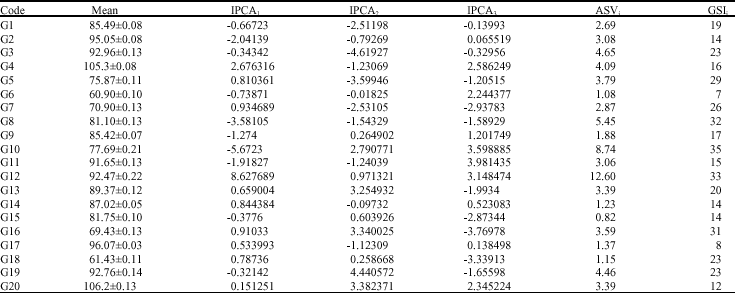
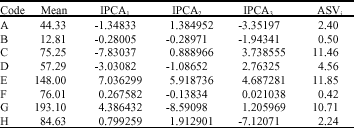
| Table 4: | First, second and third IPCA scores, ASVi and GSIi of genotypes investigated |
 | |
| Table 5: | First, second and third IPCA scores and ASVi of environments |
 | |
A large contribution of environment indicated that environments were diverse, with large differences among environmental means causing most of the variation in grain yield. The magnitude of the GE interaction was 2.5 times larger than that for genotypes, exhibiting that there were sustainable differences in the genotypic response across environments.
The results of AMMI analysis also showed that the first interaction principal component axis (IPC1) captured 36.5 of the interaction SS, IPC2 and IPC3 explained 24.5 and 22% of GE interaction SS, respectively. The mean square for IPC1 and IPC2 were significant (p<1%) and cumulatively accounted for 61% of total GE interaction. Therefore, the post-dictive evaluation using an F-test at p = 1% suggested that two IPC1 and IPC2 were significant for the model with 48 df.
In general, AMMI2 model contained 83% of the treatment SS, while the residual contained 17%. These results indicate that the AMMI model fit the data well and justifies the use of AMMI2.
IPCAs crossover (qualitative) and non-cross over interaction (quantitative): IPCA scores of genotype and environment took both positive and negative values (Table 4, 5). Consequently, a genotype that has large positive IPCA score with some environment most have large negative interactions with some other environments.
Thus, these scores presented a disproportionate genotype response (Yan and Hunt, 2001; Mohammadi et al., 2007), which was the major source of variation for any crossover interaction.
This disproportionate genotype response is referred to as crossover GE interaction for convenience. Diversely, scores with the same sign or near zero represent a non-crossover GE interaction or a proportionate genotype response (Mohammadi et al., 2007; Mohammadi and Amri, 2008).
AMMI Stability Value (ASVi): In fact ASV is the distance from zero in a two dimensional scattergram of IPCA1 (interaction principal component analysis axis 1) scores against IPCA2 scores. Since the IPCA1 score contributes more to GE sum of scores, it has to be weighted by the proportional difference between IPCA1 and IPCA2 scores to compensate for the relative contribution of IPCA1 and IPCA2 total GE sum of squares. The distance from zero is then determined using the theorem of Pythagoras (Purchase et al., 2000).
In ASV method, a genotype with least ASV score is the most stable, accordingly genotype G15, followed by G6, G18, G14, G17 and G9 were the most stable, while genotypes G3,G4, G8, G10, G12 and G19 were undesirable. Biplot analysis and ordination techniques revealed high significant differences for IPCA1, IPCA2 and IPCA3.
First three interaction principal components (IPC1, IPC2 and IPC3) explained 83% of variability in the GE interaction. Corrected grain yield can be obtained by AMMI1, AMMI2 and AMMI3 for each environment and used as a selection criteria in breeding programs. In general the importance of AMMI model is in reduction of the noise even if principal component do not cover much of the GESS (Gauch and Zobel, 1989; Gauch, 1992). Biplot analysis (Fig. 1) displayed that genotype G10 and G12 and environment C, E and G have the greatest effect in the GE interaction. Genotype number G12 has specific adaptation with environment G and E, while genotype number G10 has specific adaptability with environment C and genotype 8 has specific adaptation with environment A and D. the accessions G2 and G11 have negative GE interaction.
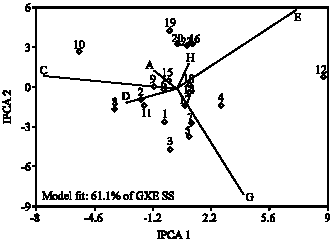 | |
| Fig. 1: | Biplot analysis of GE interaction based on AMMI2 model for the first two interaction principal component scores |
Genotypes toward the center of biplot have zero interaction, therefore have general adaptation with different mean grain yield. Genotypes G6, G9, G14, G15, G17 and G18 are located in this category and as the entries G6, G14 and G17 have mean yield over the grand mean, therefore they can be considered as stable with high performance.
Biplot of IPC1 and IPC2 (Fig. 1) covers 61.1% of GE interaction. The accessions G1, G3, G4, G5, G7, G8, G10, G16, G19 and G20 are located around the center of biplot indicating the variable of these entries (Manrique and Hermann, 2000). Genotype farther from the center of biplot show specific adaptation. The genotype G12 has specific adaptation with environment E because their angle is less than 90% and their GE interaction is positive. Genotype G1, G3, G5 and G7 have positive interaction with environment G, but as the length of the vector for genotype 3 is more on the environment G, hence it has specific adaptability with environment G. The lines G2 and G11 have positive interaction with environment D and the entry G10 show specific adaptation with environment C.
As AMMI2 has the least RMSPD (root mean square predictive difference), therefore recommendation must be based on this model (Crossa et al., 1991; Wade et al., 1995; Farshadfar and Sutka, 2006).
In pattern analysis genotypes are judged in grouping form and therefore save time and precision in interpretation and selection (Wade et al., 1995; Alagarswamy and Chandra, 1998; Farshadfar and Sutka, 2003).
Genotype Selection Index (GSI): Stability per se should however not be the only parameter for selection, because the most stable genotypes would not necessarily give the best yield performance (Mohammadi et al., 2007), hence there is a need for approaches that incorporate both mean grain yield and stability in a single criteria thats, why Kang (1991, 1993) introduced three selection criteria for simultaneous selection of yield and stability entitled: rank-sum (RSM), modified rank-sum (MRSM) and the statistics yield-stability (Ysi).
In this regard, as ASV takes into account both IPCA1 and IPCA2 that justify most of the variation of GE interaction, therefore the rank of ASVi and mean grain yield (R![]() ) are incorporated in a single selection index namely Genotype Selection Index (GSI). The least GSI is considered as the most stable with high grain yield. Based on the GSI the most desirable genotype for selection of both stability and high grain yield is G6 followed by G17 which is in agreement with the result of biplot.
) are incorporated in a single selection index namely Genotype Selection Index (GSI). The least GSI is considered as the most stable with high grain yield. Based on the GSI the most desirable genotype for selection of both stability and high grain yield is G6 followed by G17 which is in agreement with the result of biplot.
REFERENCES
- Alagarswamy, G. and S. Chandra, 1998. Parttern analysis of international sorghum multi-environment trials for grain-yield adaptation. Theor. Applied Gent., 96: 397-405.
CrossRef - Asenjo, C.A., R. Benzus and H. Acciaresi, 2003. Genotype-environment interactions in rice (Oriza sativa L.) in temperate region using the joint regression analysis and AMMI methods. Cereal. Res. Commun., 31: 97-104.
Direct Link - Basford, K.E., W.T. Federer and I.H. Delacy, 2004. Mixed model formulations for Multi-environment trails. Agron. J., 96: 143-147.
Direct Link - Crossa, J., P.N. Fox, W.H. Pfeiffer, S. Rajaram and H.G. Gauch Jr., 1991. AMMI adjustment for statistical analysis of an international wheat yield trial. Theoret. Applied Genet., 81: 27-37.
CrossRefDirect Link - Eberhart, S.A. and W.A. Russell, 1966. Stability parameters for comparing varieties. Crop Sci., 6: 36-40.
CrossRefDirect Link - Farshadfar, E., M. farshadfar and J. Sutka, 1999. Genetic analysis of phenotypic stability parameters in wheat. Acta Agron. Hung., 47: 109-116.
Direct Link - Farshadfar, E. and J. Sutka, 2003. Locating QTLs controlling adaptation in wheat using AMMI model. Cereal Res. Commun., 31: 249-254.
Direct Link - Farshadfar, E. and J. Sutka, 2006. Biplot analysis of genotype-environment interactin in durum wheat using the AMMI model. Acta Agron. Hung., 54: 459-467.
Direct Link - Finlay, K.W. and G.N. Wilkinson, 1963. The analysis of adaptation in a plant-breeding programme. Aust. J. Agric. Res., 14: 742-754.
CrossRefDirect Link - Gabriel, K.R., 1971. The biplot graphic display of matrices with application to principal component analysis. Biometrika, 58: 453-467.
Direct Link - Gauch Jr., H.G. and R.W. Zobel, 1989. Accuracy and selection success in yield trials analysis. Theor. Applied Genet., 77: 443-481.
CrossRefDirect Link - Gauch, Jr. H.G., 1988. Model selection and validation for yield trials with interaction. Biometrics, 44: 705-715.
Direct Link - Kang, M.S., 1991. Modified rank-sum for selecting high-yielding, stable corn genotypes. Cereal. Res. Commun., 19: 361-366.
Direct Link - Kang, M.S., 1993. Simultaneous selection for yield and stability in crop performance trials: Consequences for growers. Agron. J., 85: 754-757.
Direct Link - Manrique, K. and H. Hermann, 2000. Effect of GE interaction on root yield and beta-carotene content of selected sweet potato varieties and breeding clones. Trop. Agric., 119: 281-286.
Direct Link - Mohammadi, R. and A. Amri, 2008. Comparison of parametric and non-parametric methods for selecting stable and adapted durum wheat genotypes in variable environments. Euohytica, 159: 419-432.
CrossRef - Mohammadi, R., M. Armion, A. Shabani and A. Daryaei, 2007. Identification of stability and adaptability in advanced durum genotypes using AMMI analysis. Asian J. Plant Sci., 6: 1261-1268.
CrossRefDirect Link - Mohammadi, R., R. Haghparast, M. Aghaee and M. Rostaee, 2007. Biplot analysis of multi-environment trials on bread wheat yield under rainfed condition in Iran. Crop Res., 33: 170-178.
Direct Link - De Oliveira, E.J. and I.J. de Godoy, 2006. Pod yield stability analysis of runner peanut lines using AMMI. Crop Breed Applied Biotechnol., 6: 311-317.
Direct Link - Oritza, R., W.W. Wagorie, J. Hill, S. Chandra, S. Madsen and O. Stolen, 2001. Heritability and correlation among genotype by environment stability statistics for grain yield in bread wheat. Theor. Applied Genet., 103: 469-474.
CrossRef - Pourdad, S.S. and R. Mohammadi, 2008. Use of stability parameters for comparing safflower genotypes in multi-environment trials. Asian J. Plant Sci., 7: 100-104.
CrossRefDirect Link - Purchase, J.L., H. Hatting and C.S. van Denventer, 2000. Genotype x environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. South Afr. J. Plant Soil, 17: 101-107.
CrossRef - Sharma, R.C., G. Ortiz-Ferrara, J. Crossa, M.R. Bhatta and M.A. Sufian et al., 2007. Wheat grain yield and stability assessed through regional trials in the Eastern Gangetic Plains of South Asia. Euphityca, 157: 457-464.
CrossRefDirect Link - Snedecor, G.W. and W.G. Cochran, 1989. Statistical Methods. 8th Edn., Iowa State University Press, Ames, IA, USA, ISBN-13: 978-0813815619, Pages: 524.
Direct Link - Solomon, A., M. Nigussie and H. Zelleke, 2008. Genotype-environment interactionand stability analysis for grain yield of maize (zea mays L.) in Ethiopia. Asian J. Plant Sci., 7: 163-169.
Direct Link - Thamson, W.E. and S.B. Philips, 2006. Methods to evaluate wheat cultivar testing environment and improve cultivar selection protocols. Field Crops Res., 99: 87-95.
Direct Link - Vergas, M., J. Crossa., F.V. Eeuwijk, K.D. Sayre and M.P. Reynolds, 2001. Interpreting treatment×environment interaction in agronomy trials. Agron. J., 93: 949-960.
Direct Link - Yan, W., L.A. Hunt, Q. Sheng and Z. Szlavnics, 2000. Cultivar evaluation and mega-environment investigation based on the GGE biplot. Crop Sci., 40: 597-605.
CrossRefDirect Link








