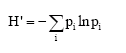Research Article
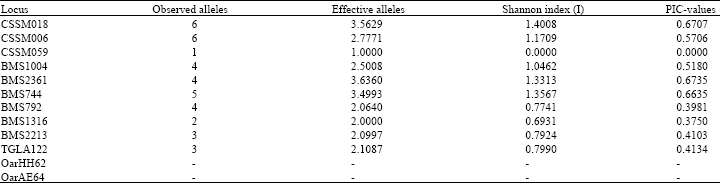
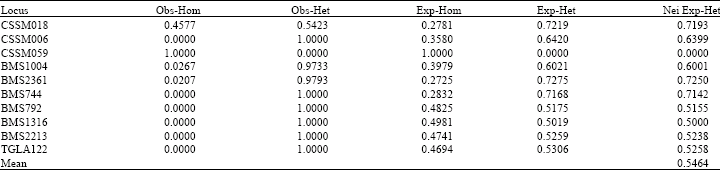
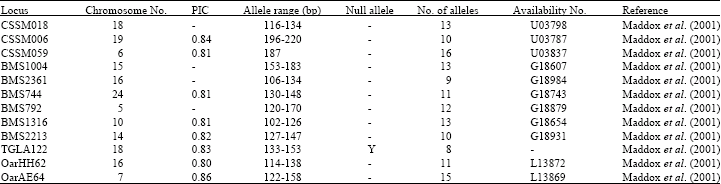
Microsatellite Variation in One Breed of Iranian Sheep with 12 Markers
University of Tarbiyat Modares, Tebran, Iran
S. Esmaeelkhanian
Department of Biotechnology, Animal Science Research Institution of Iran, Tehran, Iran
R. Vaez Torshizi
University of Tarbiyat Modares, Tebran, Iran