ABSTRACT
Nine Egyptian cultivars of Vicia faba were analyzed using electrophoretic and quantitative data from seed albumins and globulins and 100 seeds weight to measure genetic variation among faba bean cultivars in Egypt. Wide genetic variation was indicated for all the traits studied. The elctrophoregrams showed identity profile for each cutivar supporting the validity of electrophoresis of seed protein components in cultivar identification and assessing genetic variation in Vicia faba and other out-breeding plants at the infra-specific level. Negative correlation was indicated between seed globulins and both seed albumins and seed weight. Princpal component analysis and cluster analyses indicated higher role of seed albumins over seed globulins in genetic variation within Vicia faba in Egypt.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/pjbs.2007.4204.4210
URL: https://scialert.net/abstract/?doi=pjbs.2007.4204.4210
INTRODUCTION
Seed storage proteins represent a major component of human protein intake, either directly or indirectly via their consumption by livestock. Whereas cereal grains contain less than 15% protein, legume seed generally contains more than 20% (Payne, 1983; Signor et al., 2005). On a world basis, legume seeds are the second most important protein source after cereals (Perrot, 1995; Signor et al., 2005).
Characterization of the genetic variation in the available germplasm is important for further improvement of crop yield and to impart resistance to biotic and abiotic stresses (Kour and Singh, 2004). Using the morphological data in cultivar identification, studying genetic diversity and phenetic classification of plants that was previously scored for many plants (Bult and Kiang, 1992; Zviniene and Pank, 1996) is very time consuming and may be unreliable (Ahmad et al., 1997) since some of these characteristics are strongly affected by the environment. These problems have now been overcome, in many crops, by the use of electrophoresis to discriminate cultivars on the basis of genotype-specific protein markers (Barratt, 1980; Stegemann, 1983; Hussain et al., 1986). In addition, evidences provided by the separation of seed protein components in polyacrylamide gels are mostly informative at the species and infra-specific levels (Nei et al., 1978; Ladizinsky and Hymowitz, 1979; Cooke, 1984; Badr, 1995). Seed proteins data have therefore been applied to study genetic diversity at infraspecific levels (Signor et al., 2005; Mustafa et al., 2006; Sammour et al., 2007) and also for cultivar identification (Cooke, 1984; Sammour, 1988, 1990b, 1992; Krochko and Bewley, 2000; Thanh et al., 2006). In addition to morphological and seed proteins electrophoretic profiles, molecular, isozymes and immunological data were utilized in studying genetic diversity, cultivar identification and assessment of taxonomic and genetic relationships in Vicia and many other plants at generic, specific and infra-specific levels (Torres et al., 1993; Conney et al., 1998; Croft et al., 1999; Shiran and Raina, 2001; Badr et al., 2002; Zeid, 2003; Sammour, 2005; Mustafa et al., 2005, 2006; Shiran et al., 2006; Sorkheh et al., 2007). Among the mentioned techniques, the seed protein electrophoresis is considered the cheapest and less time consuming.
Using the electrophoretic techniques to study the Egyptian crop cultivars will allow us to identify between cultivars and screen the purity of the ever expanding number of cultivars, to clarify taxonomic and evolutionary problems, to exploit the important traits of landraces and wild relatives and to evaluate the genetic variation among the collected accessions. All these information are important for crop improvement programs, certification authorities and also in genetic resource management (Forde and Gardiner, 1986; Gardiner and Forde, 1988; Sammour, 1990b; Kour and Singh, 2004).
Using electrophoresis of the total seed proteins showed some shortage in discrimination between Vicia faba cultivars (Gardiner and Forde, 1988; Thanh et al., 2006). In this study, the author tried to use electrophoretic data of both seed albumins and globulins in addition to some other traits to study the genetic variations among nine Egyptian cultivars of Vicia faba, to test the validity of the applied technique for faba bean cultivar identification and also as a basic requirement for further crop traits improvement programs.
MATERIALS AND METHODS
This study was conducted at Botany Department, Faculty of Science, Tanta University at 2005-2006. Seeds of Vicia faba L. cultivars were collected from the Agricultural research station, institute of legumes, Sakha, Kafr El-Sheikh, Egypt. The cultivars list is shown in Table 1.
Quantitative estimation of seed proteins: Albumin and Globulin proteins were extracted successively from 20 mg air-dried defatted seed meals in 1000 μL extraction solution (distilled water for albumins then 5% NaCl w/v for globulins) for 24 h at -4°C, according to Croy and Gatehouse (1985) with some modifications. The extracts were centrifuged for 10 min at 10000 g and quantitative estimation of total proteins was made according to Bradford (1976). Three replicas were made for each sample.
SDS-PAGE: For electrophoresis, extracts were mixed with 10 % sucrose and used directly for electrophoresis or kept at -20°C until use. Dissociating polyacryamide gel electrophoresis (SDS-PAGE) was adopted after Laemmli (1970) with some modifications. Samples of 15 μL were applied onto main gels of 17% acrylamide concentration using bromophenol blue as a front dye. Electrophoresis was carried out using 25 mM tris/glycine buffer pH 8.3. The gel was stained overnight by slow shaking in 100 mL staining mixture (0.05% Coammassie Brillient Blue-R250 in 50% methanol, 7% glacial acetic acid and 43% distilled water) and de-stained by slow shaking in the stain solvent. The subunit molecular weight of the protein bands was determined according to Weber and Osberne (1969) through a standard curve established for each gel. Each band was considered as a character for which the presence or absence was coded by 1 or 0, respectively in a data matrix for statistical analysis. Density of the protein bands was not considered.
100 seeds-weight: Weight 100 seed in each sample and notes it; the difference in weight present among samples may indicate the genetic variation.
Data analysis: The genetic diversity among the populations of the studied cultivars (evaluated by Dice similarity index) and multivariate analysis (factor analyses and cluster analysis) were made using the software SPSS for windows package (Version 10) copyright (C) 1999, SPSS INC. A dendrogram was constructed through the complete linkage-joining rule. Nei's similarity index (Nei and Li, 1979) was calculated also among the studied cultivars where:
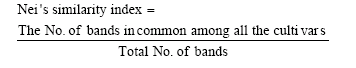 |
The similarity matrix values were converted into percentages.
RESULTS
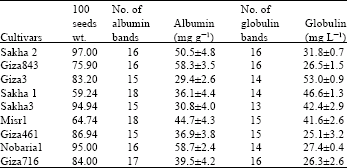
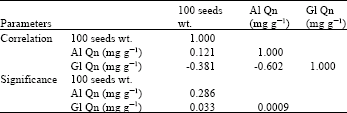
The results obtained in this study showed little variation in the number of bands of either globulins [13-16 bands] or albumins electrophoregrams [15-18 bands] (Fig. 1, Table 1). The studied cultivars showed higher Nei's similarity index for globulins (57.79%) than that for albumins electrophoregrams (42.857). The studied cultivars of Vicia faba exhibited wide genetic variation in relation to each of 100 seeds weight, albumin and globulin seed protein contents (Table 1). A negative correlation was exhibited between seed globulin content and both seed weight (-0.381) (significant) and seed albumin content (-0.602) (highly significant), while 100 seeds weight showed small positive non-significant correlation (0.121) with seed albumin content (Table 2).
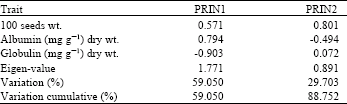
The first two components account for 88.752% of the total variance of the studied. Separate percentages of variation attributed to the first two components are 59.05 and 29.703%, with Eigen values of 1.771 and 0.891, respectively.
| Table 1: | Hundred seeds weight, No. of bands in the electrophoregrams and seed contents of Albumin and globulin in faba bean cultivars |
 | |
| Table 2: | Correlation coefficients among seed weight, seed globulin and seed albumin contents of faba bean cultivars |
 | |
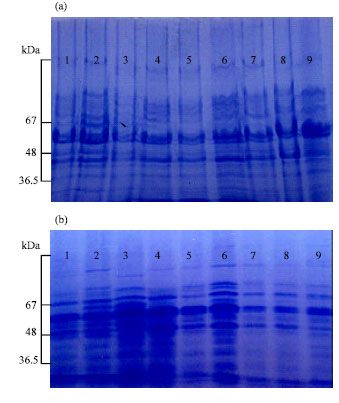 | |
| Fig. 1: | Electrophoregrams of seed proteins (a) albumins and (b) globulins of Vicia faba cultivars: 1 (sakha2), 2 (Giza843), 3 (Giza3), 4 (sakha1), 5 (sakha3), 6 (misr1), 7 (Giza461), 8 (nobaria1), 9 (Giza716) |
Albumin content showed higher coefficient in the first component (PRIN1), while the 100 seeds wt. showed higher variation coefficient with the second component (PRIN2) (Table 3). The third component showed significant sharing of variation to associate with globulin content (not shown data). In this study, the first two component, were considered.
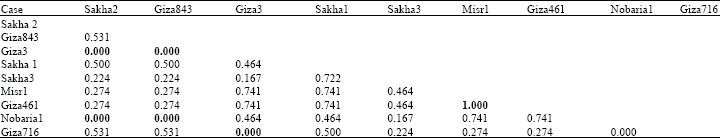
Cluster analysis of electrophoretic globulin data (Table 4) showed highest genetic similarity between misr 1 and giza 461 according to Dice measure. Farthest genetic distance was exhibited between giza 3 and each of sakha 2, giza 843 and giza 716 and between nobaria 1 and both sakha 2 and giza 843. However, highest genetic similarity index was exhibited between misr 1 and giza 461.
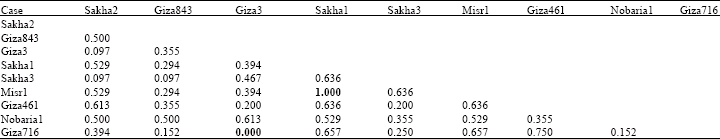
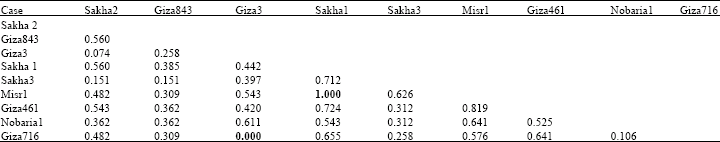
Based on electrophoretic data of albumin alone or aggregated with globulin data, nearest genetic distance was exhibited between sakha1 and misr 1, while farthest genetic distance was exhibited between giza3 and giza 716 according to Dice measure (Table 5, 6). Dendrograms based on cluster analysis according to Dice measure (Fig. 2-4) showed two major groups. In case of electrophoretic data of albumin alone or aggregated with globulin data, the smaller group included three cultivars (Fig. 2, 4), while in case of electrophoretic data of globulin alone, the smaller group included four cultivars. The studied cultivars were distributed into three groups at a genetic distance of 0.21, based on albumin electrophoretic data (Fig. 2) and at a genetic distance of 0.19, based on globulin electrophoretic data (Fig. 3).
| Table 3: | Principal component analysis of Vicia faba cultivars, Eigen-values and percent of variation accounted by the first two components |
 | |
| Table 4: | Genetic similarity index (Dice measure) among Vicia faba cultivars based on globulin electrophoretic data |
 | |
| Table 5: | Genetic similarity index (Dice measure) among Vicia faba cultivars based on albumin electrophoretic data |
 | |
| Table 6: | Genetic similarity index (Dice measure) among Vicia faba cultivars based on albumin and globulin electrophoretic data |
 | |
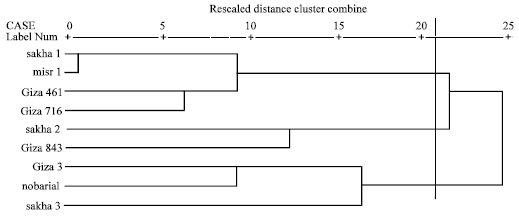 | |
| Fig. 2: | Dendrograms based on electrophoretic data of seed albumins according to Dice |
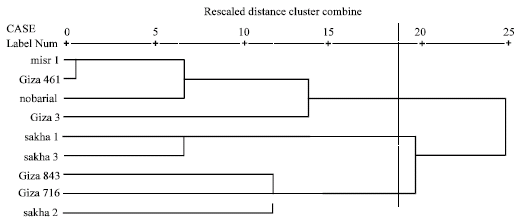 | |
| Fig. 3: | Dendrograms based on electrophoretic data of seed globulins according to Dice |
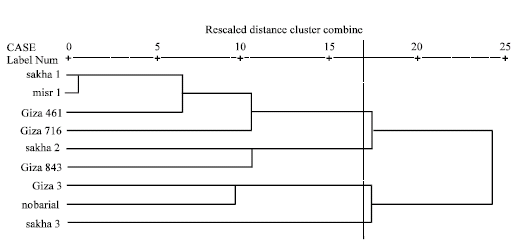 | |
| Fig. 4: | Dendrograms based on electrophoretic data of seed albumins and globulins according to Dice |
On aggregating electrophoretic data of albumins and globulins, an arbitrary genetic distance of 0.17 was used to divide the studied cultivars into four groups (Fig. 4).
DISCUSSION
This study was conducted to study the genetic variations among the Egyptian cultivars of Vicia faba as a basic requirement for further crop traits improvement programs and to test the validity of the applied technique for faba bean cultivar identification. In the present investigation, high genetic variation was observed for the amount of albumin and globulin proteins of the seed meal and 100-seed weight. Also, in spite of the narrow variation in the number of bands in both albumins and globulins electrophoregrams, the Nei's similarity coefficient among all the studied cultivars was small (57.79% for globulins and 42.857 for albumins), indicating a wide genetic variation in both seed albumins and globulins of the studied faba bean cultivars. This genetic variation may be indicated in the polymorphism exhibited in the minor bands and in the major bands intensities, which gave each cultivar its specific electrophoregram for either albumins or globulins. These identity electrophoregrams of the studied cultivars can be used as passport data for their genetic identity and can be used as a good tool for testing core collection concepts and organizing genetic diversity in Vicia faba. These results support the validity of seed protein electrophoresis as a powerful tool for cultivar identification, clarifying taxonomic and evolutionary problems and studying genetic diversity (Ladizinsky and Hymowitz, 1979; Cooke, 1984; Sammour, 1988, 1990a,b; 1992; Krochko and Bewley, 2000; Signor et al., 2005; Mustafa et al., 2006; Thanh et al., 2006; Sammour et al., 2007). Also, the wide genetic variation observed for seed albumins and globulins and seed weight, which is a genetic trait (Rees, 1997; Sammour et al., 2007), indicated that improvement through simple selection for these traits is possible. However, broadening the genetic base from diverse sources is recommended to include most of the genetic determinants of these traits (Laghetti et al., 1998; Ghafoor et al., 2001; Sammour et al., 2007).
The association between globulins and both seed weight and albumins in the seed of the studied cultivars exhibited negative values which was significant in case of seed weight and highly significant in case of seed albumins. This suggests some kind of association in the genetic control of these traits, which may be supported by the high percentage of variance (88.752% of the total variance) accounted for by the first two principal components in multivariate analysis. These results may be in contradiction with other previous views (Polignano et al., 1979; Dixit et al., 1995; Granati et al., 2003). This contradiction is probably because the author used the seed protein components, albumins and globulins, rather than the total seed proteins. Thanh et al. (2006) concluded that using protein components (Albumins, globulins, prolamins and glutelins) in cultivar identification is recommended to expose the minor genetic variation between the cultivars of a specific crop. This was applied previously for Lathyrus sp. (Przybylska et al., 1999) and Phaseolus lunatus L. (Vargas et al., 2000). The conclusion of Thanh et al. (2006) may be supported by the present principal component analysis, where the PRIN1 accounted for 59.05% of the total variance with the highest load for seed albumins (0.794), which indicates crucial role of seed albumin variation in genetic variation and discriminating the faba bean cultivars. Further research is needed to determine exactly the genetic control of this correlation. These facts call for more large collections to be analyzed and for more extensive research work to ascertain the contents of the protein components in the seeds and their relationships with other seed traits.
The superior role of seed albumins over seed globulins in cultivar discrimination and genetic variation in Vicia faba at the infra-specific level is indicated in the results of cluster analysis of electrophoretic data that showed similarity in genetic relationships among the studied cultivars and the topology of the dendrograms in case of seed albumin alone and aggregated with seed globulin data. This could be obviously noted in grouping the same three cultivars, Giza3, Nobaria1 and Sakha3 in one group in the dendrograms based on albumin alone or aggregated with globulin.
Nadal et al. (2007) reported that Natural out-crossing is the major cause of loss of varietal purity in faba bean (Vicia faba L.) affecting both the regeneration-multiplication of germplasm collections and the production of certified or basic seed by plant breeders. Faba bean is a partially cross-pollinated species with a rate of out-crossing ranging from 4 to 84% (Bond and Poulsen, 1983). The results obtained here supports the validity of seed proteins electrophoresis for cultivar identification, studying genetic diversity and taxonomic relationships of Vicia faba and also other out-breeding plants at the intra-specific level.
REFERENCES
- Ahmad, M., A.G. Fautrier, D.J. Burritt and D.L. McNeil, 1997. Genetic diversity and relationships in Lens species and their F1 interspecific hybrids as determined by SDS-PAGE. N.Z. J. Crop Hortic. Sci., 25: 99-108.
CrossRefDirect Link - Badr, A., 1995. Electrophoretic studies of seed proteins in relation to chromosomal criteria and the relationships of some taxa of Trifolium. Taxon, 44: 183-191.
Direct Link - Badr, A., H. Sayed-Ahmed, L.E. Watson and A. El-Shanshouri, 2002. Ancestors of white clover (Trifolium repens L.), as revealed by isozyme polymorphisms. Theor. Appl. Genet., 106: 143-148.
CrossRefPubMedDirect Link - Barratt, D.H., 1980. Cultivar identification of Vicia faba (L.) by sodium dodecyl sulphate polyacrylamide gel electrophorsis of seed globulins. J. Sci. Food Agric., 31: 813-819.
CrossRefDirect Link - Bradford, M.M., 1976. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal. Biochem., 72: 248-254.
CrossRefPubMedDirect Link - Bult, C.J. and Y.T. Kiang, 1992. Electrophoretic and morphological variation within and among natural populations of the wild soybean, Glycine soja. Bot. Bull. Acad. Sin., 33: 111-112.
Direct Link - Conny, B., A. Smussen and A., Aron, 1998. Chloroplast DNA characters, phylogeny and classification of Lathyrus (Fabaceae). Am. J. Bot., 85: 387-401.
Direct Link - Croft, A.M., E.C.K. Pang and P.W.J. Taylor, 1999. Molecular analysis of Lathyrus sativus L. (grasspea) and related Lathyrus species. Euphytica, 107: 167-176.
CrossRefDirect Link - Gardiner, S.E. and M.B. Forde, 1988. Identification of cultivars and species of pasture legumes by sodium dodecylsulphate polyacrylamide gel electrophoresis of seed proteins. Plant Varieties Seeds, 1: 13-16.
Direct Link - Ghafoor, A., A. Sharif, Z. Ahmad, M.A. Zahid and M.A. Rabbani, 2001. Genetic diversity in black gram (Vigna mungo L. Hepper). Field Crop Res., 69: 183-190.
CrossRefDirect Link - Granati, E., V. Bisignano, D. Chiaretti, P. Crino and G.B. Polignano, 2003. Characterization of Italian and exotic Lathyrus germplasm for quality traits. Genet. Resour. Crop Evol., 50: 273-280.
CrossRef - Krochko, J.E. and J.D. Bewley, 2000. Seed storage proteins in cultivars and subspecies of alfafa (Medicago sativa). Seed Sci. Res., 10: 423-434.
CrossRefDirect Link - Ladizinsky, G. and T. Hymowitz, 1979. Seed protein electrophoresis in taxonomic and evolutionary studies. Theor. Applied Genet., 54: 145-151.
CrossRefDirect Link - Laemmli, U.K., 1970. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature, 227: 680-685.
CrossRefDirect Link - Laghetti, G., B.L. Pienaar, S. Padulosi and P. Perrino, 1998. Ecogeographical distribution of Vigna savi in Southern Africa and some areas of the Mediterranean basin. Plant Genet. Resourc. Newslett., 115: 6-12.
Direct Link - Mustafa, A.Z.M.A., A. Badr, M.A. El-Galaly, A.A. Mobarak and M.G. Hassan, 2005. Genetic diversity among Mentha populations in Egypt as reflected by isozyme polymorphism. Int. J. Bot., 1: 188-195.
CrossRefDirect Link - Mustafa, A.M.A., A. Badr, M.A. El-Galaly, A. Mobarak and M.G. Hasan, 2006. Genetic diversity among Ocimum populations in Egypt as reflected by morphological, seed proteins and isozyme polymorphism. Int. J. Bot., 2: 261-269.
CrossRefDirect Link - Nadal, S., M.J. Suso, M.T. Moreno and J.I. Cubero, 2007. Evaluating the effectiveness of small isolation distances for the field multiplication of Vicia faba L. using a morphological recessive marker Plant Genet. Resour. Newslett., 136: 11-13.
Direct Link - Nei, M., P.A. Fuerst and R. Chakraborty, 1978. Subunit molecular weight and genetic variability of proteins in natural populations. Proc Natl. Acad. Sci., USA., 75: 3359-3362.
PubMedDirect Link - Nei, M. and W.H. Li, 1979. Mathematical model for studying genetic variation in terms of restriction endonucleases. Proc. Natl. Acad. Sci. USA., 76: 5269-5273.
CrossRefPubMedDirect Link - Przyblska, J., Z. Zimniak-Przybylska and P. Krajewski, 1999. Diversity of seed albumins in some Lathyrus species related to L. sativus L.: An electrophoretic study. Gen. Resour. Crop Evol., 46: 261-266.
Direct Link - Sammour, R.H., 2005. Molecular reassessment of relationships within Vicieae using electrophoretic and immunochemical techniques. Afr. Crop Sci. J., 13: 27-39.
Direct Link - Shiran, B. and S.N. Raina, 2001. Evidence of rapid evolution and incipient speciation in Vicia sativa species complex based on nuclear and organellar RFLPs and PCR analysis. Genet. Resour. Crop Evol., 48: 519-532.
CrossRefDirect Link - Shiran, B., A. Roghaieh, M. Shahram and A.M. Reza, 2006. Potential use of random amplified polymorphic DNA marker in assessment of genetic diversity and identification of rapeseed (Brassica napus L.) cultivars. Biotechnology, 5: 153-159.
CrossRefDirect Link - Signor, C.L., K. Gallardo, J.M. Prosperi, C. Salon, L. Quillien, R. Thompson and G. Duc, 2005. Genetic diversity for seed protein composition in Medicago truncatula. Plant Genet. Resour., 3: 59-71.
Direct Link - Sorkheh, K., B. Shiran, T.M. Gradziel, B.K. Epperson, P.M. Gomez and E. Asadi, 2007. Amplified fragment length polymorphism as a tool for molecular characterization of almond germplasm: Genetic diversity among cultivated genotypes and related wild species of almond and its relationships with agronomic traits. Euphytica, 156: 327-344.
CrossRefDirect Link - Thanh, V.C., P.V. Phuong, P.H.H. Uyen and P.P. Hien, 2006. Application of protein electrophoresis SDS-PAGE to evaluate genetic purity and diversity of several varieties. Proceedings of the International Workshop on Biotechnology in Agriculture, October 20-21, 2006, Nong Lam University Ho Chi Minh City, pp: 192-194.
- Torres, A.M., N.F. Weeden and A. Martin, 1993. Linkage among isozyme, RFLP and RAPD markers in Vicia faba. Theor. Applied Genet., 85: 937-945.
Direct Link








