Research Article
Study on Variation of Potato Varieties Using Electrophoretic Tuber Storage Proteins
University of Zanjan, Zanjan, Iran
Lamei Javad
Agricultural Research Center of Khirabad, Zanjan, Iran
Potato (Solanum tuberosum L.) is the world’s fourth most important crop after rice, wheat and corn and its importance is growing (Mullins et al., 2006). The potato tuber is also an important source of industrial starch. Potato juice is a waste product from industrial starch manufacture and contains approximately 1.5% (w/v) of soluble protein. Osborne and Campbell (1896) reported that the major protein in potato tubers was a globulin which they termed, tuberin,. More recently, Racusen and Foote (1980) reported that a glycoprotein of Mr about 45000 accounted for about 20% of the total soluble protein in potato and proposed the alternative name patatin, based on patata which is the original American Indian-derived Spanish word for potato (Shewry, 2003). Potato juice protein has been classified into three major groups: patatins, protease inhibitors and other proteins. Patatins and protease inhibitors, each comprising approximately 40% of total tuber protein, are mostly considered as storage proteins, in addition to their lipolytic and inhibitory activities, respectively (Jorgensen et al., 2006).
The genus Solanum consists of about 900 species, 235 of which form tubers (Hawkets, 1990). However, the ability to form tubers is not an essential requirement for the functional value of a species in potato breeding. The potato therefore, is a crop plant with an enormously wide gene pool (Menke et al., 1996).
The spread of the potato from its centre of origin in the high Andes of South America to other parts of the globe and the historical consequences of this have been well documented (Messer, 2000; Shewry, 2003; Bauw et al., 2006; Jorgensen et al., 2006). The potato production area is occurred in western part of Iran, including Azerbaijan, Ardabil and Zanjan provinces. Classification and organization of genotypes into systematic relationships is based on a wide array of morphological, biochemical and molecular descriptors. To measure genetic relationships accurately, the ideal descriptor would provide an unbiased estimate of total genome variation and be sufficiently abundant to minimize errors due to sampling variance. Potato tuber proteins have been studied and their high quality verified by several investigators (Nakasone et al., 1972; Stegeman et al., 1973; Kapoor et al., 1975; Racusen and Foote, 1980; Rajapakse and Imai, 1991; Satu et al., 2005; Bauw et al., 2006; Jorgensen et al., 2006). High genetic variability in protein content exists in cultivars of sweet potato (Walter et al., 1984; Babu et al., 1990; Woolfe, 1992; Barta et al., 2003). Due to the diversity of tuber proteins within the group tuberosum, protein analysis by electrophoresis is a convenient and effective method of studying genetic variation (Andrews, 1987). The same technique can be used to monitor genetic stability in gene banks where large quantities of plant materials are conserved. In such analysis, the test environment should be standardized, thus allowing genetic variation to appear clearly (Rajapakse and Imai, 1991; Barta et al., 2003). Electrophoretic patterns of soluble proteins and isozymes have been used as a powerful tool for the study of genetic variability of Solanum species since the sixties of the last century. In Europe, collections of potato cultivars were discriminated by their protein and esterase patterns (Barta et al., 2003).
Tubers are suitable for separating proteins, since they represent a stable source of proteins and provide a uniform material for protein extraction. The protein patterns of potato were found to be stable regardless of the addition of growth regulators or changes in environmental factors (Rajapakse and Imai, 1991). For any breeding programs, specially for hybridization, it is important to have an understanding of the affinity among different varieties and cultivars. Genetic fingerprinting has been accomplished traditionally through the use of isozymes, total seed proteins, tuber storage protein and more recently through various types of molecular markers (Alvarez et al., 2003; Barta et al., 2003; Singh et al., 2006; Lara et al., 2006). The aim of this study is used of potato storage protein patterns for evaluation of genetic variation on potato varieties.
Plant materials: Thirteen varieties of potato cultivated in potato breeding farm in Khirabad Agricultural Research Station during the 2004 (Table 1). This site was located about 30 km east of the Zanjan province in Iran. Same field-grown tubers of varieties were harvested after maturity and used for protein extraction.
Protein extraction: About 1 g from each varieties tuber (same weight) was comminuted with 1 mL sample buffer (Laemli, 1970). Sap samples were prepared by pressing small cubes of tuber using a hand-managed press (garlic press) at 4°C. The liquid portion was separated by centrifugation at 4000 g for 10 min at 4°C.
The supernatant was centrifuged again and the resulting supernatant kept for 3 h at room temperature and either used directly for electrophoresis without heat treatment or kept at -21°C for up to one month.
Preparation of slab gel: The gels were prepared according to the method described by Laemmli (Laemmli, 1970). A vertical slab gel 1 mm thick consisted of a running gel (about 100 mm high), layered with a stacking gel (about 12 mm high). Stock solutions were mixed in the following way, to make one volume of running gel: 0.33 volumes of acryl amide solution containing 300 g L-1 acryl amide and 8 g L-1 N, N-methylene-bis-acrylamide (Bis), 0.2 volumes of tris (hydroxymethel)-aminomethane and hydrochloric acid (Tris-HCL, 1.5 M, pH. 8.9) and 0.003 volumes of freshly prepared 100 g L-1 ammonium persulfate (AMPS) were mixed with 0.47 volumes of deionized water (final concentration: acrylamid: 10 %: Bis 0.27%). The mixture was degassed prior to the addition of 0.0003 volumes of N, N, N, N-tetramethyl ethylenediamine (TEMED) and poured between glass plates (140x140 mm). The mixture was layered with 0.375 M Tris-HCL (pH. 8.9) and allowed to polymerize for 30-40 min. When the separation line was visible the overlying layer was changed once and the gel was allowed to stand overnight or the stacking gel was prepared immediately. The stacking gel mixture was prepared as follows: 0.2 volume of acrylamide solution containing 200 g L-1 acrylamide and 10 g L-1 Bis, 0.25 volume of 0.5 M Tris-HCL (pH. 6.7), 0.004 volume of 100 g L-1 AMPS were mixed with 0.5 volumes of deionized water. The mixture was degassed, 0.001 volume of TEMED were added and poured over the separating gel after removing the overlying layer of Tris-HCL solution. A Teflon comb was fixed on top of the stacking gel which was allowed to polymerize for 2 h. The comb had been removed and protein samples were loaded at 20 g L-1 per lane. One molecular weight marker lane were maintained for identification. The migrating position of these markers is indicated in Fig. 1.
| Table 1: | Potato varieties used for protein electrophoresis |
 | |
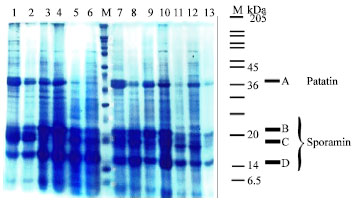 | |
| Fig. 1: | Comparison of SDS-PAGE protein banding patterns of 13 varieties. Lane 1-13: Protein pattern, Lane M: Molecular weight of marker protein-subunits is indicated in kDa (6.5, 14, 20, 24, 29, 36, 45, 55, 66, 84, 97, 116, 205). A, B, C, D: 4 major soluble tuber proteins. A is patatin and B, C, D are sporamin isomers |
Electrophoresis: The glass plate with the gel were fixed in the electrophoretic apparatus filled with running buffer (0.025 M Tris, 0.192 M glycine). Electrophoresis was done at a constant current of 15 mA per plate for 7 h. Staining was done by coomassie blue (coomassie brillant blue R 250, SIGMA Co.,USA) as 2 g L-1 coomassie blue that dissolved in ethanol: acetic acid: water = 45: 10: 45 at room temperature for 5 h. The de-staining was done using ethanol: acetic acid: water (25: 10: 65) at room temperature. Only clear and unambiguous bands were recorded based on their relative mobility. Binary data matrix was generated taking 1 as presence and 0 as absence. The statistical calculation were done using STATISTICA program.
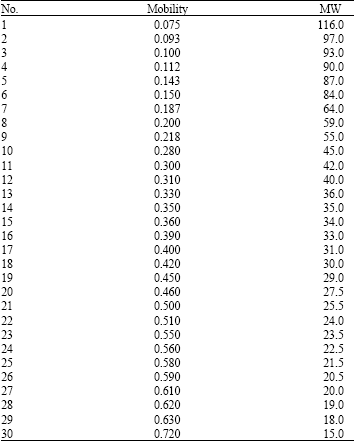
The electrophoretic tuber storage protein banding of 13 varieties of potato has shown in Fig. 1. The protein profile revealed a total of 30 electrophoretic bands (Table 2). None of the varieties showed identical banding pattern. The maximum number of bands present in any species was 19 in Finna (1, 2, 4, 6, 7, 9, 10, 12, 13, 14, 16, 18, 19, 20, 23, 25, 27, 29, 30) and the minimum number of bands were found 8 in (6, 12, 17, 20, 24, 17, 19, 30). Several specific bands were also observed such as band number 5 in Picasso, 21 in Marifona; 22 in Ditta. All the varieties had five bands including 6, 12, 19, 29 and 30. The mobility and molecular weight of 30 protein bands is shown in Table 2.
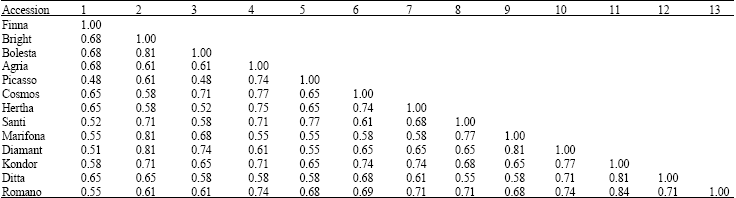
Binary data matrix was generated taking 1 as presence and 0 as absence. The statistical calculation were done using STATISTICA program. The similarity matrix showed the maximum similarity is occurred with Romano and Kondor (84%) (Table 3). They were clustered together between around 0.16 to 0.52 level of distance, with a mean value of 0.34, showing that the genetic dissimilarity between potato varieties is smaller than their similarity (1-0.34 = 0.66). Cluster analysis revealed that these 13 varieties of potato could be grouped in 4 classes (Fig. 2): 1) Finna 2) Bolesta, Marifona, Diamant and Bright 3) Agria, Cosmos, Herta, Romano, Kondor and Ditta 4) Santi and Picasso. ad2ad4**ad5
Since the some morphological characteristics such as color of skin tubers and flowers were studied, consequently the Kondor and Romano had purple color in tubers skin. In addition to this skin color, three varieties had pink flowers. These including Kondor, Romano and Diamant. The color of flowers in others is white.
In this study we obtained 4 major bands (A, B, C, D) in the 14-45 kDa. Molecular weight range. According to Fig. 1. Three observed major bands (B, C, D) were those of sporamin isomers. They could be seen as two or three separated major bands towards the middle portion of the gel in both native and SDS-containing gels.
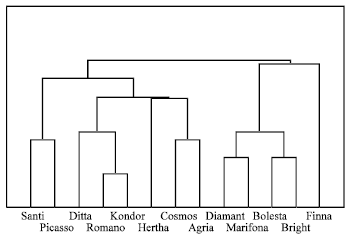 | |
| Fig. 2: | Dendrogram based on cluster analysis of potato tuber proteins (SDS-PAGE). Cluster analysis (0.34 level of distance) revealed that these 13 varieties of potato grouped in 4 classes (1) Finna (2) Bolesta, Marifona, Diamant and Bright (3) Agria, Cosmos, Herta, Romano, Kondor and Ditta and (4) Santi and Picasso |
| Table 2: | Electrophoretic characteristics of 30 protein bands in the 15-116 kDa MW |
 | |
Together they accounted for 60 to 70 percent of the soluble proteins extracted from fresh tissue (Babu et al., 1990). The molecular weight fell between 14-20 kDa for these bands, confirming their identity as sporamin isomers (Li and Oba, 1985; Babu, 1996).
| Table 3: | Similarity matrix based on protein banding pattern in 13 varieties of potato |
 | |
Another major protein which could be easily detected in SDS-PAGE had a molecular weight of approximately 40 kDa and was identified as patatin (Rajapakse and Imai, 1991; Shewry, 2003; Revina et al., 2004; Bauw et al., 2006; Jorgensen et al., 2006).
For an effective breeding program, information concerning the extent and nature of genetic diversity within a crop species is essential. It is particularly useful for characterizing individual accessions and cultivars and as a general guide in the selection of the parents for hybridization. Genetic fingerprinting has been accomplished traditionally through the use of isozymes, total seed proteins, tuber storage proteins and more recently through various types of molecular markers (Hamrich and Godt, 1997; Alvarez et al., 2003; Barta et al., 2003; Lara et al., 2006; Marfil et al., 2006; Pierre et al., 2006; Singh et al., 2006; Zosimo et al., 2006). Although, DNA-based markers provide powerful tools for discerning variations within crop germplasm and for studying evolutionary relationships, but the capacity of protein markers is yet high and effective, specially in undevelopment countries. Based on the results of this research, we can confirm that cultivar differences in protein polymorphism can be revealed by applied electrophoretic patterns of storage protein via SDS-PAGE in potato cultivars.