Research Article
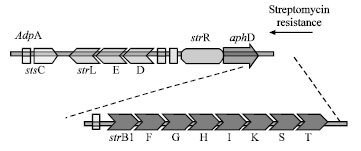
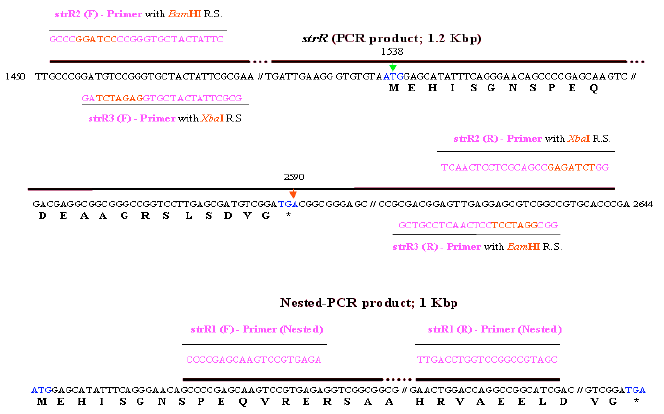
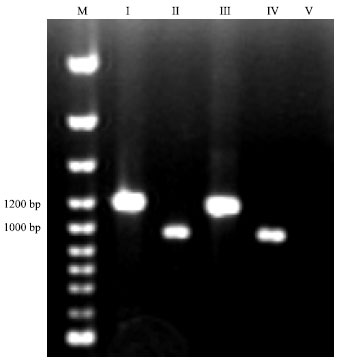
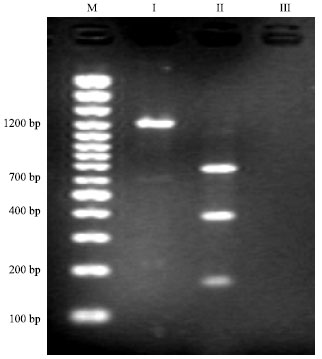
Detection of Cloned strR, an Antibiotic Regulatory Gene, using RFLP and Nested PCR
Department of Biology, Faculty of Sciences, Isfahan University, Isfahan, Islamic Republic of Iran
Majid Motovali-Bashi
Department of Biology, Faculty of Sciences, Isfahan University, Isfahan, Islamic Republic of Iran
Naser Golbang
Department of Biology, Faculty of Sciences, Isfahan University, Isfahan, Islamic Republic of Iran
Farshad Darvishi
Department of Biology, Faculty of Sciences, Isfahan University, Isfahan, Islamic Republic of Iran