Research Article
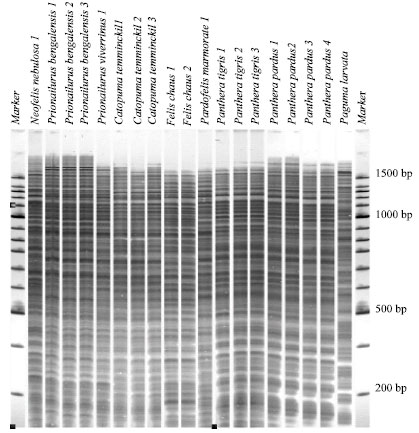
Genetic Relationships among Wild Felidae in Thailand Using AFLP Markers
Department of Biochemistry
Arunrat Chaveerach
Department of Biology, Faculty of Science, Khan Kaen University, Khan Kaen, 40002, Thailand
Suporn Nuchadomrong
Department of Biochemistry
Nison Sattayasai
Department of Biochemistry
Prapansak Chaveerach
Department of Veterinary Public Health, Faculty of Veterinary Medicine, Khan Kaen University, Khan Kaen, 40002, Thailand
Alongkoad Tanomtong
Department of Biology, Faculty of Science, Khan Kaen University, Khan Kaen, 40002, Thailand
Krit Pinthong
Biology Program, Faculty of Science and Technology, Surin Rajabhat University, Surin, 32000, Thailand