Research Article
Molecular Modelling Analysis of the Metabolism of Naproxen
School of Biomedical Sciences, Faculty of Health Sciences, The University of Sydney, P.O. Box 170, Lidcombe, NSW 1 825, Australia
Naproxen (S)-6-methoxy-α-methyl-2-naphthaleneacetic acid, [NAP] is a non-steroidal anti-inflammatory drug (NSAID) that has been widely used in the treatment of rheumatoid arthritis and osteoarthritis (Konstantianos et al., 1993). The drug is almost fully absorbed when given orally in daily doses of 500-1500 mg. Although NAP is effective and considered to be safe, it has a number of side-effects including gastrointestinal toxicity (Aabakken et al., 1989), nephrotoxicity (Cox et al., 1990), jaundice (Bass, 1974; Law and Knight, 1976). It has been suggested that the hypersensitive response of the drug is due to hepatitic injury caused by the drug. NAP has also been found to induce lipid peroxidation in isolated rat hepatocytes (Yokoyama et al., 1994). This is actually believed to be caused by the Reactive Oxygen Species (ROS) generated during NAP metabolism. It is possible that electrophilic metabolites from NAP may cause depletion of cellular glutathione, thus compromising the antioxidant status of the cell and hence producing oxidative stress.
Although the predominant species found in the plasma is NAP (Calvo et al., 1987), that is extensively bound to plasma proteins mainly albumin (Bischer et al., 1995), it is excreted almost exclusively in the form of metabolites in the urine (Anderson and Hansen, 1992). Conjugation with glucuronic acid is the major metabolic pathway for NAP, as it is true for many other drugs containing carboxylic acids. Thus the major metabolite of NAP in humans is naproxen-β-1-O-acyl glucuronide (NAP-AGLU). Other important metabolic pathways include O-dealkylation to form 6-O-desmethylnaproxen (DNAP), conjugation with glycine to form naproxen-glycine conjugate (NAP-GLY), conjugation with sulfate to form naproxen-sulfate [NAP-SU]. The enzymes involved in O-demethylation are CYP2C9 and CYP1A2. DNAP which has both carboxyl and phenolic groups can also undergo acyl glucuronidation to form DNAP acyl glucuronide (DNAP-AGLU). It can also undergo phenolic glucuronidation and sulfation (Jaggi et al., 2002; Sidelmann et al., 2001).
It has been suggested that acyl glucuronides are intrinsically more reactive, being able to undergo rearrangement, via acyl migration (catalysed by hydroxide) to form 2-, 3- and 4-O positional isomers that are resistant to hydrolysis by β-glucuronidases (Lo et al., 2001). They can also undergo other reactions including hydrolysis and adduction with biomolecules (Faed, 1984; Dickinson et al., 1993) so that when rearrangement does not occur, 1-O-acyl glucuronides (NAP-AGLU and DNAP-AGLU) may react readily with nucleophilic centres such as -H, -OH and -NH2 groups in proteins and peptides, resulting in loss of glucuronic acid moiety (Williams and Dickinson, 1994). It has been suggested that the drug-protein adducts formed may be the cause of allergy and hepatotoxicity due to NAP. Also, the depletion of cellular glutathione (GSH) may induce cellular injury by compromising the antioxidant status of the cell.
In this study, molecular modelling analyses have been carried out using the program Spartan ’02 (Spartan, 2002) to investigate the relative stability of NAP and its metabolites leading to a better understanding of the toxicity of NAP and its metabolites.
Computational Methods
The geometries of NAP and its metabolites, DNAP, NAP-AGLU, NAP-GLY, NAP-SU and DNAP-AGLU have been optimized based on molecular mechanics (Fig. 1), semi-empirical and DFT calculations, using the molecular modelling program Spartan ’02. Molecular mechanics calculations were carried out using MMFF force field. Semi-empirical calculations were carried out using the routine PM3. The order of calculations: molecular mechanics followed by semi-empirical followed by DFT assisted in reaching global minimum. It was found that when semi-empirical calculations were not preceded by molecular mechanics calculations, structure could be embedded in a local minimum. DFT calculations were carried using the program Spartan ’02 at B3LYP/6-31G* level.
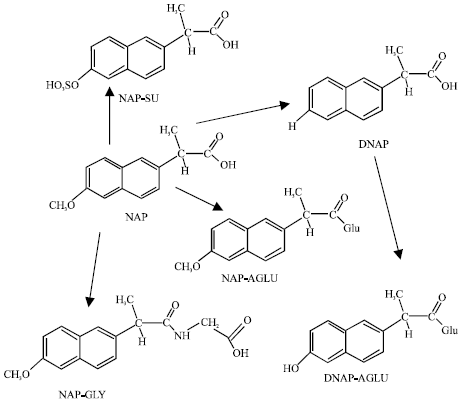 | |
| Fig. 1: | Metabolic pathways for NAP (Adapted from Sidelmann et al., 2001). |
In optimization calculations, a RMS gradient of 0.001 was set as the terminating condition. Although RMS value gradient of 0.001 will not be sufficiently low for vibrational analysis, it is believed to be sufficient for calculations associated with electron energy levels. For the optimized structures, single point calculations were carried to give heat of formation, enthalpy, entropy, free energy, dipole moment, solvation energy, energies for HOMO and LUMO. The gradient in single point calculation as a fraction was found to be less than 0.001. The study was carried out in the School of Biomedical Sciences, The University of Sydney during January to May 2006.
Table 1 gives the total energy, heat of formation as per PM3 calculation, enthalpy, entropy, free energy, dipole moment, energies of HOMO and LUMO as per both PM3 and DFT calculations for NAP, DNAP, NAP-AGLU, NAP-GLY, NAP-SU and DNAP-AGLU. Fig. 2-7 give the regions of negative electrostatic potential (greyish-white envelopes) in (a), HOMOs (where red indicates HOMOs with high electron density) in (b), LUMOs in (c) and density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) in (d) as applied to the optimized structures of NAP and its metabolites, DNAP, NAP-AGLU, NAP-GLY, NAP-SU and DNAP-AGLU.
The calculated solvation energies of NAP and its metabolites, DNAP, NAP-AGLU, NAP-GLY, NAP-SU and DNAP-AGLU from PM3 calculations in kcal mol-1 are, respectively -12.26, -15.10, -17.93, -52.42 and -55.79 and their dipole moments from DFT calculations are 6.4, 6.0, 4.6, 7.1 and 6.9, respectively. The high solvation energy and dipole moment values indicate that NAP and its metabolites would all be soluble in water. The terminal metabolites NAP-AGLU and DNAP-AGLU are expected to be much more soluble in water than NAP and its other metabolites.
| Table 1: | Calculated thermodynamic and other parameters for naproxen and its metabolites [DM stands for dipole moment] |
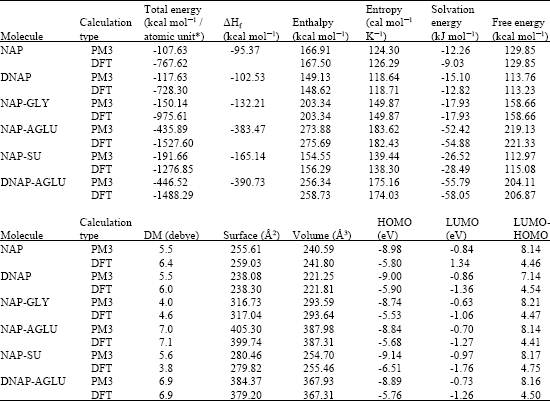 | |
| * in atomic units from DFT calculations | |
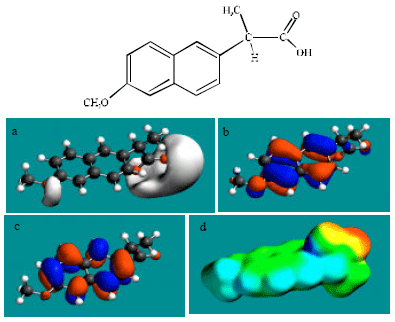 | |
| Fig. 2: | Structure of NAP giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface electric (where red indicates negative, blue indicates positive and green indicates neutral) |
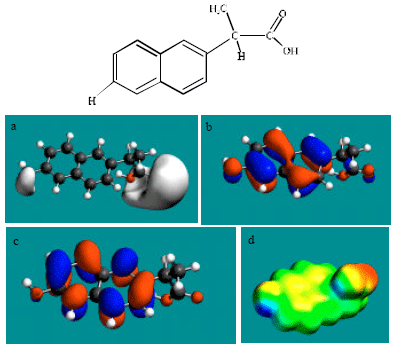 | |
| Fig. 3: | Structure of DNAP giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface electric (where red indicates negative, blue indicates positive and green indicates neutral) |
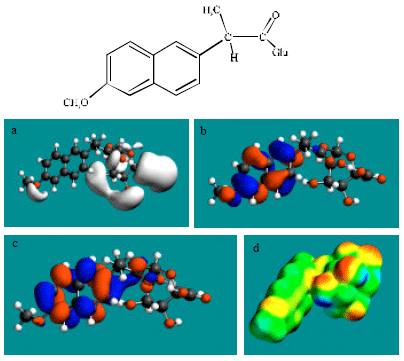 | |
| Fig. 4: | Structure of NAP-AGLU giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface electric (where red indicates negative, blue indicates positive and green indicates neutral) |
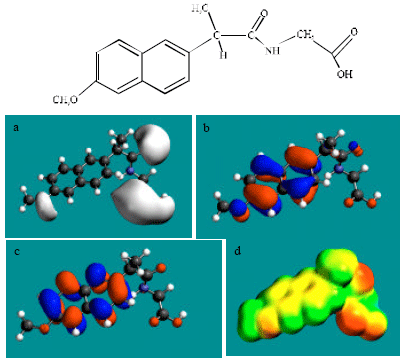 | |
| Fig. 5: | Structure of NAP-GLY giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface electric (where red indicates negative, blue indicates positive and green indicates neutral) |
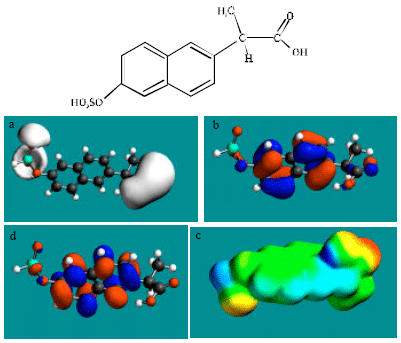 | |
| Fig. 6: | Structure of NAP-SU giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface electric (where red indicates negative, blue indicates positive and green indicates neutral) |
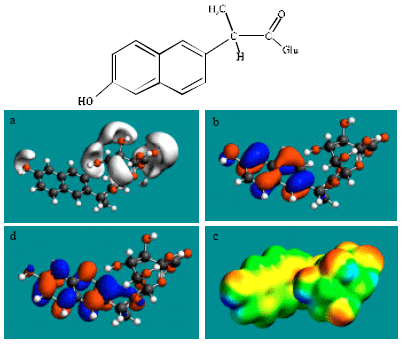 | |
| Fig. 7: | Structure of DNAP-AGLU giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) density of electrostatic potential on the molecular surface electric (where red indicates negative, blue indicates positive and green indicates neutral) |
NAP and its metabolites are found to have LUMO-HOMO energy differences ranging from 4.4 to 4.5 eV from DFT calculations, indicating that in terms of kinetic lability they would be similar (neither highly inert nor exceedingly labile).
In the case of NAP, electrostatic potential is found to be more negative around carboxyl and methoxy oxygen atoms, indicating that the positions may be subject to electrophilic attack. In the case of DNAP and NAP-AGLU, electrostatic potential is found to be more negative around carboxyl and hydroxyl oxygen atoms, indicating that the positions may be subject to electrophilic attack. In the case of NAP-GLY, electrostatic potential is found to be more negative around carboxyl, carbonyl and methoxy oxygen atoms, indicating that the positions may be subject to electrophilic attack. In the case of NAP-SU, electrostatic potential is found to be more negative around carboxyl and sulfate oxygen atoms, indicating that the positions may be subject to electrophilic attack. In the case of NAP-GLY, electrostatic potential is found to be more negative around carboxyl, carbonyl and methoxy oxygen atoms, indicating that the positions may be subject to electrophilic attack. In the case of DNAP-AGLU, electrostatic potential is found to be more negative around carboxyl and hydroxyl oxygen atoms, indicating that the positions may be subject to electrophilic attack.
In the case of NAP, DNAP, NAP-AGLU, NAP-GLY, NAP-SU and DNAP-AGLU, the HOMOs with high electron density and LUMOs are found to be centred on the non-hydrogen atoms of the two fused phenyl rings. The convergence or close proximity of some HOMOs with high electron density with those of negative electrostatic potential give further support to the idea that the positions may indeed be subject to electrostatic attack.
As stated earlier Fig. 2-5d and 7d give the density of electrostatic potential on the molecular surfaces of NAP and its metabolites DNAP, NAP-AGLU, NAP-GLY, NAP-SU and DNAP-AGLU where red indicates electron-rich regions, green indicates neutral and blue indicates electron-deficient regions. It can be seen that the surfaces of NAP and the metabolite NAP-SU abound most in electron-deficient regions. This means that NAP and NAP-SU may be most subject to nucleophilic attack e.g., that by cellular glutathione leading its depletion, thus compromising the antioxidant status of the cell and hence causing cellular toxicity. However, as noted earlier, neither NAP nor any of its metabolites is expected to be highly labile so that the rate of reaction of NAP and NAP-SU with glutathione may be low. Other metabolites DNAP, NAP-AGLU and DNAP-AGLU also cannot be excluded from consideration of reaction with cellular glutathione and other nucleophiles as they also have some electron-deficient regions. However, as in the case of NAP and NAP-SU, the rates of such reactions are expected to be low.
Although naproxen is considered to be a safe non-steroidal anti-inflammatory drug, it has a number of side-effects including gastrointestinal toxicity, nephrotoxicity, jaundice and hepatotoxicity. Molecular modelling analyses based on molecular mechanics, semi-empirical and DFT (at B3LYP/6-31G* level) calculations show that there are both electron-rich and electron-deficient regions on the surfaces of NAP and its metabolites so that they may be subject to both electrophilic and nucleophilic attacks. The latter means that they can react with cellular glutathione, causing its depletion and thus inducing cellular toxicity. However, the rates of such reactions are expected to be low as neither NAP nor any of its metabolites is likely to be highly labile.
| Abbreviations | |
| NAP | Naproxen, [(S)-6-methoxy-α-methyl-2-naphthaleneacetic acid |
| NSAID | Non-steroidal anti-inflammatory drug |
| ROS | Reactive oxygen species |
| DNAP | O-dealkylation to form 6-O-desmethylnaproxen |
| NAP-AGLU | Naproxen-β-1-O-acyl glucuronide |
| NAP-SU Naproxen-sulfate | |
| NAP-GLY | Naproxen-glycine conjugate |
| NAP-AGLU | Naproxen-β-1-O-acyl glucuronide |
| GSH | Reduced form of glutathione |
| GSCOC | Glutathione carbonyl chloride |
| LUMO | Lowest unoccupied molecular orbital |
| HOMO | Highest occupied molecular orbital |
| DFT | Density functional theory |