Research Article
Molecular Modelling Analysis of the Metabolism of Zebularine
School of Biomedical Sciences, Faculty of Health Sciences, The University of Sydney, Australia
Zebularine (ZEB) is the generic name for 2(1H)-pyrimidinone riboside, a pyrimidinone ribinucleoside that targets epigenetic modulation of DNA methylation (Cheng et al., 2003; Yoo et al., 2004). More precisely, it is a potent inhibitor of DNA methyltransferases (Zhou et al., 2002) that are a class of enzymes involved in the epigenetic silencing of tumour suppressor genes (Yoo et al., 2004). It was originally synthesized as an inhibitor of cytidine deaminase (EC 3.54.5, CDA) (McCormack et al., 1980). ZEB is quite stable with half-lives of 44 and 68 h at pH 1.0 and 2.0, respectively and shows no evidence of decomposition after more than a week at pH 5. At pH 7.4, the half-life is 508 h (Marquez et al., 2003). It has been suggested that the oral activity of ZEB, as applied to the reactivation of silenced p16 expression, is due to its enhanced chemical stability (Cheng et al., 2003). As a CDA inhibitor, ZEB is found to have a Ki value of 2 μM, which represents a 10-fold reduction in potency relative to the less stable prototypic inhibitor tetrahydrouridine (THU) (McCormack et al., 1980). The inhibition of DNA methylation by ZEB is believed to result from the formation of a covalent adduct between the enzyme and ZEB-substituted DNA (Ben-Kasus et al., 2005). It has been hypothesized the metabolic activation of ZEB requires it to be phosphorylated and incorporated into DNA. It forms a deoxynucleotide triphosphate that incorporates into DNA at the GCGC target site of DNA methyl transferase. The resulting complex functions as a mechanism-based inhibitor of DNA methylation (Ben-Kasus et al., 2005).
Based on both in vitro and in vivo activity in mammalian cells resulting from the reactivation of a dormant p16 tumour suppressor gene (Cheng et al., 2003), ZEB has been proposed for clinical evaluation as an oral antitumour agent (Klecker et al., 2006). ZEB was also found to possess modest antitumour activity against murine B16 melanoma, P388 leukaemia and L1210 leukaemia and showed activity against when administered i.p. or orally (Driscoll et al., 1991). Furthermore, ZEB enhances the activity of decitabine which is a clinically used DNA methyl transferase inhibitor in both human and murine leukaemia cell lines (Beumer et al., 2006).
It should be noted that ZEB has been found to have poor oral bioavailability in rhesus monkey, Fisher rat and CD2F1 mouse (Holleran et al., 2005) which may be due to its rapid hepatitic metabolism to uridine that is further metabolized to uracil (Klecker et al., 2006). Biochemically ZEB behaves like an analogue of cytodine but with its own unique properties (Ben-Kasus et al., 2005). The 2-(1H)-pyrimidinone ring of ZEB lacks the amino group normally found in the 4-position of the nucleobase cytosine that makes it susceptible to nucleophilic addition of water across the 3,4-double bond to form a covalent hydrated C-4 adduct.
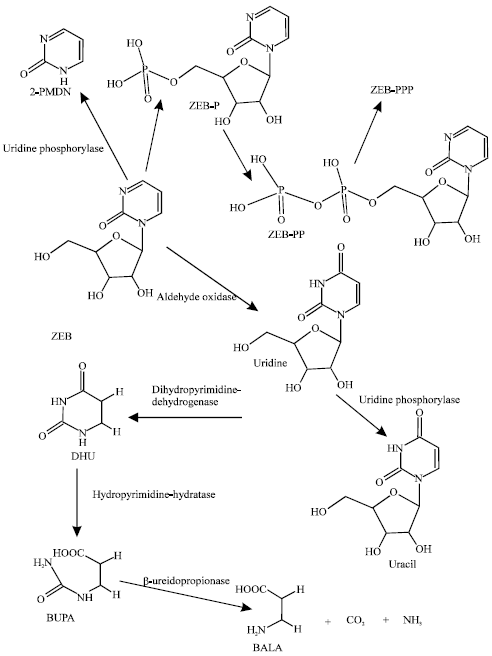 | |
| Fig. 1: | Metabolic pathways for ZEB and its metabolites (Based on Klecker et al., 2006) |
Figure 1 summarizes metabolism of ZEB. It includes: oxidation of ZEB to uridine by aldehyde hydrogenase, removal of its ribose moiety and that from uridine by uridine phosphorylase (EC 2.4.2.3) (Klecker et al., 2006) to produce 2-pyrimidinone (2-PMDN) and uracil, respectively; reduction of uracil to dihydrouracil (DHU) catalysed by dihydropyrimidine dehydrogenase (EC 1.3.1.2) and subsequent hydrolysis of DHU by hydropyrimidine hydratase to β-ureidipropionic acid (BUPA) and further hydrolysis of BUPA to carbon dioxide, ammonia and β-alanine (BALA) by β-ureidopropionase (Voet and Voet, 1995). Since incorporation into DNA appears to be a requirement for activity of ZEB, it has been suggested that ZEB must undergo phopshorylation followed by subsequent conversion to the corresponding 2’-deoxynucleotide before it can be incorporated (Ben-Kasus et al., 2005). No information is currently available on any side-effects of ZEB.
In this study, molecular modelling analyses have been carried out using the program (Spartan, 2002) to investigate the stability of ZEB and its metabolite with the aim of providing significant information on their toxicity (or lack of it). In particular, the study would show whether ZEB and its metabolites could compromise antioxidant status of the cell and/or could induce DNA damage. Modelling analyses have also been carried out of mono-, di- and tri-phosphorylated adducts of ZEB namely ZEB-P, ZEB-PP and ZEB-PPP.
Computational Methods
The geometries of ZEB and its metabolites uridine, uracil, DHU, BUPA, BALA, 2-PMDN, ZEB-P, ZEB-PP and ZEB-PPP have been optimized based on molecular mechanics, semi-empirical and DFT calculations, using the molecular modelling program Spartan ’02. Molecular mechanics calculations were carried out using MMFF force field.
Semi-empirical calculations were carried out using the routine PM3. The order of calculations: molecular mechanics followed by semi-empirical followed by DFT assisted in reaching global minimum. It was found that when semi-empirical calculations were not preceded by molecular mechanics calculations, structure could be embedded in a local minimum. DFT calculations were carried using the program Spartan ’02 at B3LYP/6-31G* level. In optimization calculations, a RMS gradient of 0.001 was set as the terminating condition. Although RMS value gradient of 0.001 will not be sufficiently low for vibrational analysis, it is believed to be sufficient for calculations associated with electron energy levels. For the optimized structures, single point calculations were carried to give heat of formation, enthalpy, entropy, free energy, dipole moment, solvation energy, energies for HOMO and LUMO. The gradient in single point calculation as a fraction was found to be less than 0.001. The study was carried out in the School of Biomedical Sciences, The University of Sydney during January to April 2006.
Table 1 gives the total energy, heat of formation as per PM3 calculation, enthalpy, entropy, free energy, dipole moment, energies of HOMO and LUMO as per both PM3 and DFT calculations for ZEB and its metabolites uridine, uracil, 2-PMDN, DHU, BUPA, BALA, ZEB-P, ZEB-PP and ZEB-PPP. Figure 2-10 give the regions of negative electrostatic potential (greyish-white envelopes) in (a), HOMOs (where red indicates HOMOs with high electron density) in (b), LUMOs in (c) and density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) in (d) as applied to the optimized structures of the optimized structures of ZEB and its metabolites uridine, uracil, 2-PMDN, DHU, BUPA, BALA, ZEB-P and ZEB-PP. The figure of ZEB-PPP has not been included.
The calculated solvation energies of ZEB and its metabolites uridine, uracil, DHU, BUPA and BALA from PM3 calculations in kcal mol-1 are, respectively -21.45, -20.02, -14.15, -14.35, -13.64, -20.35 and -14.59, respectively. The values for ZEB-P, ZEB-PP and ZEB-PPP could not be calculated.
| Table 1: | Calculated thermodynamic and other parameters of ZEB and its metabolites |
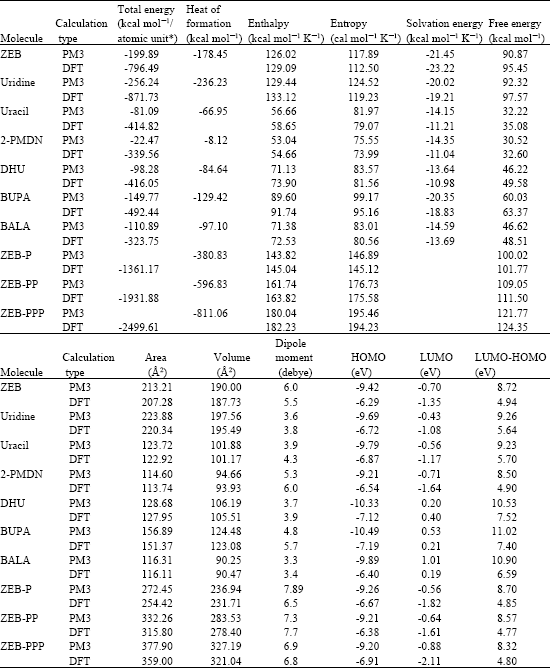 | |
| * In atomic units from DFT calculations | |
The dipole moments of ZEB, uracil, 2-PMDN, DHU, BUPA, BALA, ZEB-P, ZEB-PP and ZEB-PPP from DFT calculations are 5.5, 3.8, 4.3, 6.0, 3.9, 5.7, 3.4, 6.5, 7.7 and 6.8 indicating that ZEB and all its metabolites are highly polar. The results indicate ZEB and its metabolites would be soluble in water and therefore would be easily excreted in the urine.
ZEB and all its metabolites are found to have relatively large LUMO-HOMO energy differences indicating that they would be kinetically inert. ZEB-P, ZEB-PP, ZEB-PPP and 2-PMDN are expected to be more labile than ZEB and the other metabolites as they have somewhat lower LUMO-HOMO energy differences.
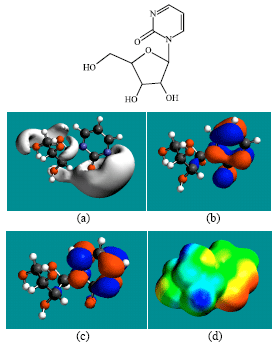 | |
| Fig. 2: | Structure of ZEB giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
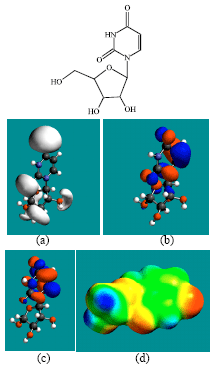 | |
| Fig. 3: | Structure of uridine giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
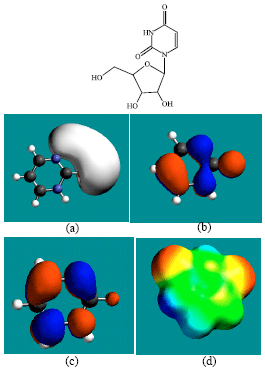 | |
| Fig. 4: | Structure of uracil giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface s (where red indicates negative, blue indicates positive and green indicates neutral) |
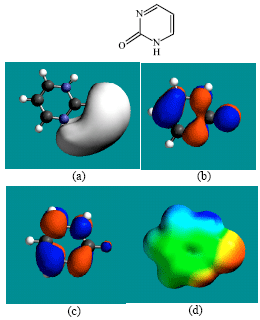 | |
| Fig. 5: | Structure of 2-PMDN giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
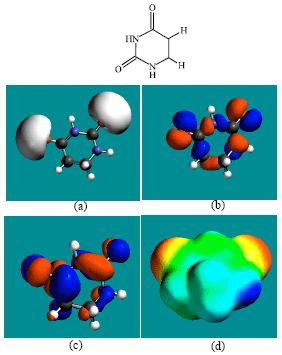 | |
| Fig. 6: | Structure of DHU giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
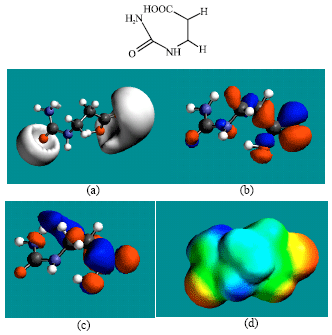 | |
| Fig. 7: | Structure of BUPA giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
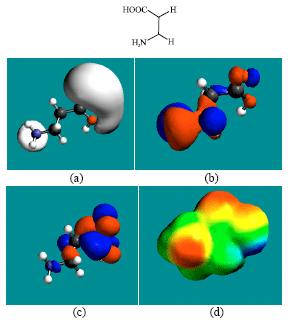 | |
| Fig. 8: | Structure of BALA giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
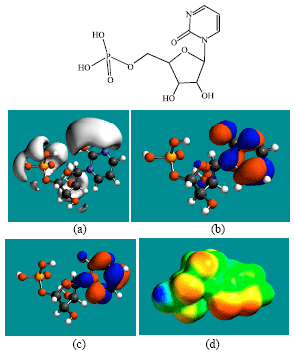 | |
| Fig. 9: | Structure of ZEB-P giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
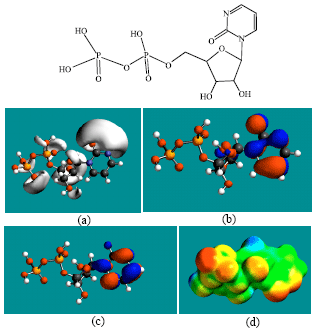 | |
| Fig. 10: | Structure of ZEB-PP giving in: (a) the electrostatic potential (greyish envelope denotes negative electrostatic potential), (b) the HOMOs, (where red indicates HOMOs with high electron density) (c) the LUMOs (where blue indicates LUMOs) and in (d) the density of electrostatic potential on the molecular surface (where red indicates negative, blue indicates positive and green indicates neutral) |
In the case of ZEB, uridine and uracil, the electrostatic potential is found to be more negative around hydroxyl and carbonyl and ethereal oxygen atoms indicating that the positions may be subject to electrophilic attack. In the case of 2-PMDN and DHU, the electrostatic potential is found to be more negative around oxygen and nitrogen atoms indicating that the positions may be subject to electrophilic attack. In the case of BUPA and BALA, the electrostatic potential is found to be more negative around carbonyl and hydroxyl oxygen atoms and the amino nitrogen atoms, indicating that the positions may be subject to electrophilic attack. In the case of ZEB-P, ZEB-PP and ZEB-PPP, the electrostatic potential is found to be more negative around hydroxyl and carbonyl and ethereal and phosphate oxygen atoms indicating that the positions may be subject to electrophilic attack This is discussed more fully later when nature of surface charge densities are considered.
In the case of ZEB, both HOMOs with high electron density and LUMOs are found to be centered on the non-hydrogen atoms of the pyrimidone ring. In the case of uridine, both HOMOs with high electron density are found to be centered on most of the non-hydrogen atoms of the pyrimidinone ring and some non-hydrogen atoms of the ribose ring whereas LUMOs are found to be centered on the non-hydrogen atoms of pyrimidinone ring.
The presence of both electron-rich (red) and electron-deficient (blue) regions on the surface of ZEB and its metabolites (Fig. 2d and 10d) indicate that they can be subject to electrophilic and nucleophilic attacks. The metabolites DHU and BUPA that have somewhat greater abundance of electron-deficient regions than ZEB and other metabolites, indicating that they may be more subject to nucleophilic attack such as that glutathione and nucleobases in DNA. Reaction with glutathione would cause glutathione depletion, thus inducing cellular toxicity and oxidation of nucleobases would cause DNA damage. However, the two metabolites are found to have the larger LUMO-HOMO energy differences so that they would be less labile. This means that the rate of such adverse reactions may not be significant.
Zebularine is a pyrimidinone ribinucleoside that is a potent inhibitor of DNA methyltransferases. Based on both in vitro and in vivo activity in mammalian cells, ZEB has been proposed for clinical evaluation as an oral antitumour agent. Molecular modelling analyses show that ZEB and all its metabolites have kinetic stability. Thus, although the presence of electron-deficient regions on the molecular surfaces of ZEB and its metabolites indicate that they may react with cellular glutathione and may cause oxidation of nucleobases in DNA, in actual fact the rates of such adverse reactions are not expected to be significant.
Abbreviations
| ZEB: | Zebularine, 2(1H)-pyrimidinone riboside |
| CDA: | Cytidine deaminase |
| DHU: | Dihydrouracil |
| THU: | Tetrahydrouridine |
| 2-PMDN: | 2-pyrimidinone |
| BUPA: | β-ureidipropionic acid |
| BALA: | β-alanine |
| LUMO: | Lowest unoccupied molecular orbital |
| HOMO: | Highest occupied molecular orbital |
Fazlul Huq is grateful to the School of Biomedical Sciences, The University of Sydney for the time release from teaching.