Research Article
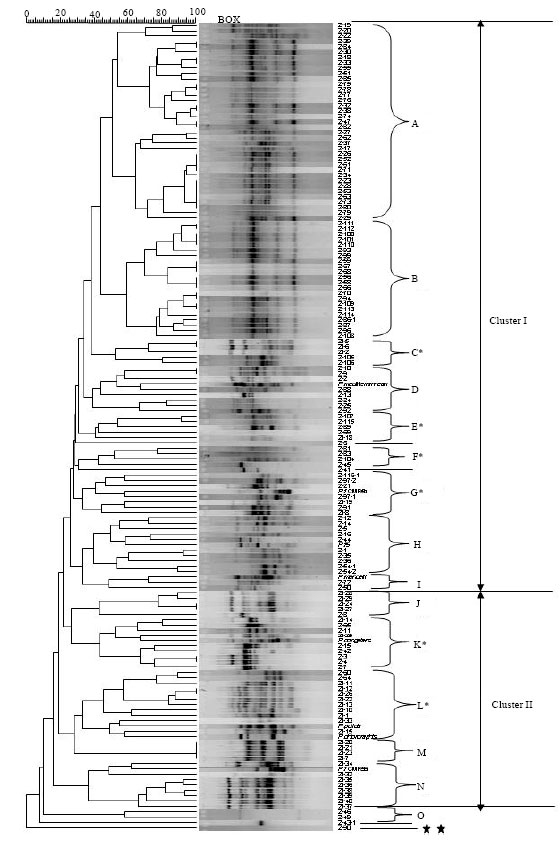
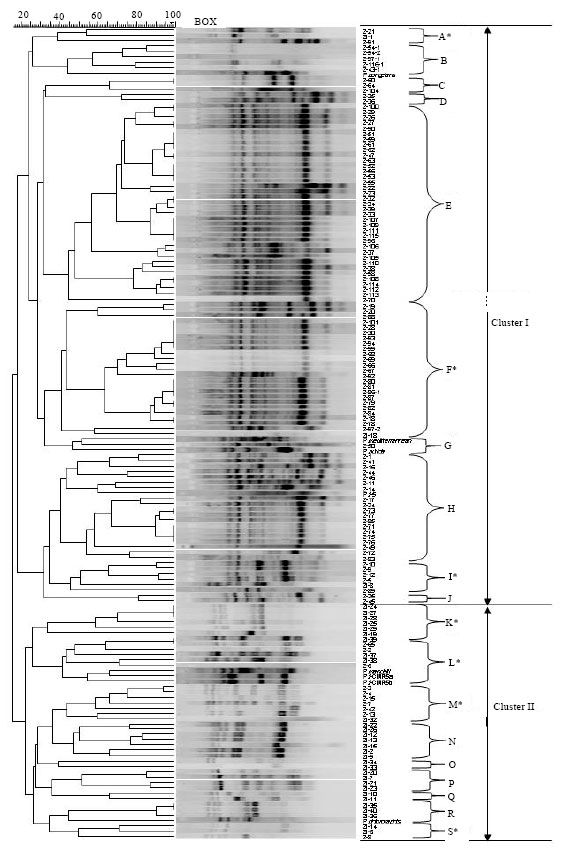
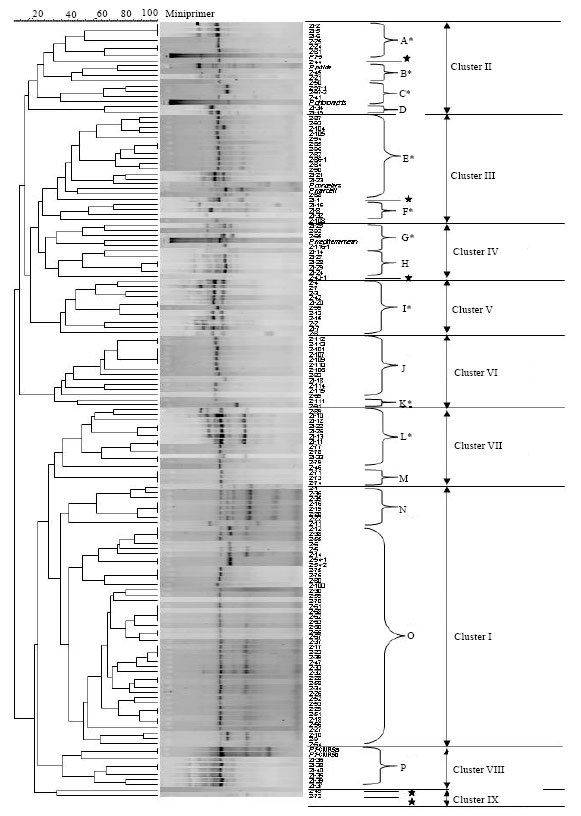
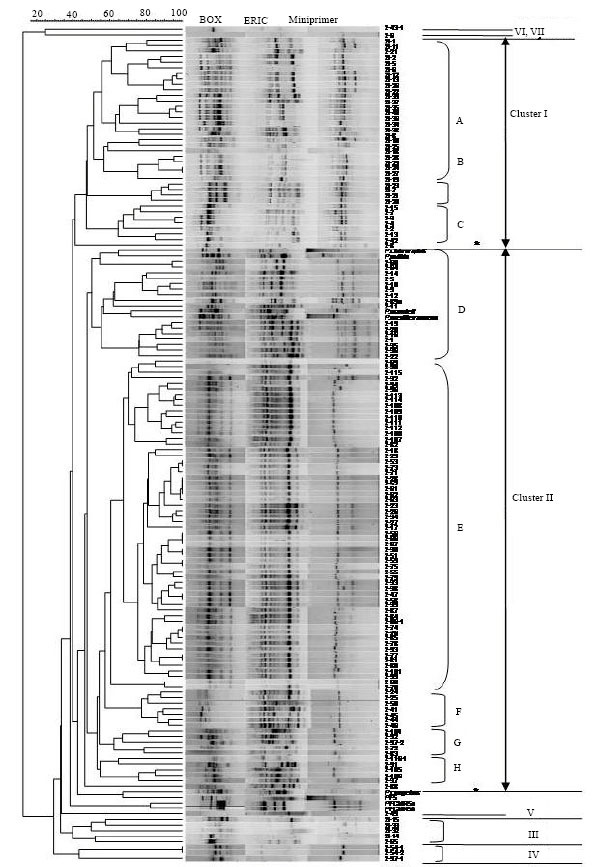
Comparison of Three Typing Methods for Evaluating the Diversity of Pseudomonas fluorescens in the Rhizosphere
Triticeae Research Institute, Sichuan Agricultural University, Yaan, Sichuan 625014, China
Pengfei Qi
Triticeae Research Institute, Sichuan Agricultural University, Yaan, Sichuan 625014, China
Renlin Xu
Agriculture and Agri-Food Canada, ECORC, Canada
James T. Tambong
Agriculture and Agri-Food Canada, ECORC, Canada
Zeinab R. Djama
Agriculture and Agri-Food Canada, ECORC, Canada
Wei Li
Triticeae Research Institute, Sichuan Agricultural University, Yaan, Sichuan 625014, China