Research Article
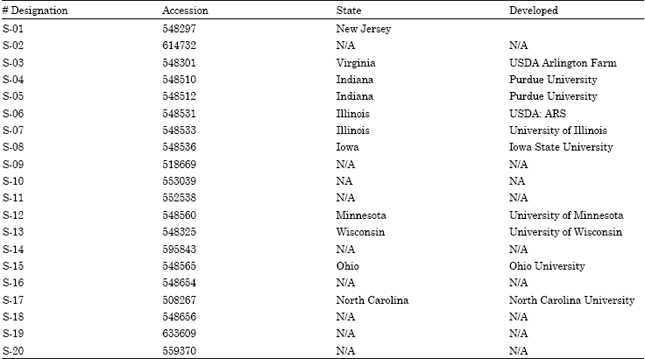
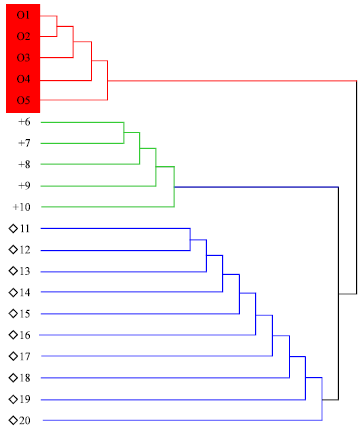
Genetic Diversity of North American Soybean (Glycine max L.) Cultivars as Revealed by RAPD Markers
Department of Environmental and Plant Biology, Ohio University Zanesville, Ohio, 43701, USA
Abdel-Salam G. Abdel Salam
Department of Statistics, Oklahoma University, Stillwater, 74078, USA