Research Article
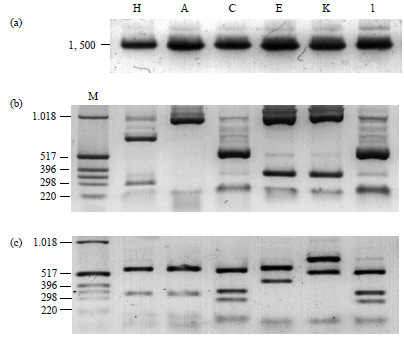
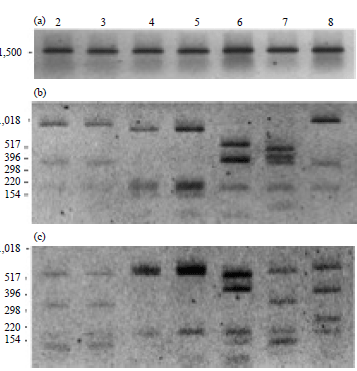
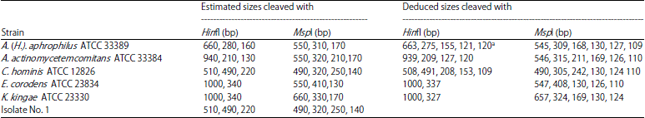
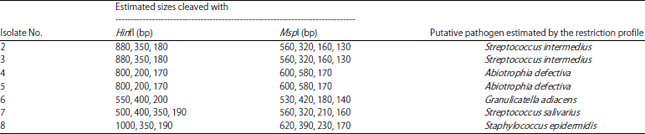
Identification of HACEK Group Bacteria from Blood Samples of Patients with Infective Endocarditis by PCR-RFLP of the 16s rRNA Gene
Division of Molecular Microbiology, Department of Microbiology, Iwate Medical University, 2-1-1 Nishitokuta Yahaba-Cho, Shiwagun, 028-3694 Iwate, Japan
Yu Shimoyama
Division of Molecular Microbiology, Department of Microbiology, Iwate Medical University, 2-1-1 Nishitokuta Yahaba-Cho, Shiwagun, 028-3694 Iwate, Japan
Yoshitoyo Kodama
Division of Molecular Microbiology, Department of Microbiology, Iwate Medical University, 2-1-1 Nishitokuta Yahaba-Cho, Shiwagun, 028-3694 Iwate, Japan
Shihoko Tajika
Division of Molecular Microbiology, Department of Microbiology, Iwate Medical University, 2-1-1 Nishitokuta Yahaba-Cho, Shiwagun, 028-3694 Iwate, Japan
Shigenobu Kimura
Division of Molecular Microbiology, Department of Microbiology, Iwate Medical University, 2-1-1 Nishitokuta Yahaba-Cho, Shiwagun, 028-3694 Iwate, Japan
Minoru Sasaki
Division of Molecular Microbiology, Department of Microbiology, Iwate Medical University, 2-1-1 Nishitokuta Yahaba-Cho, Shiwagun, 028-3694 Iwate, Japan
LiveDNA: 81.23694