Research Article
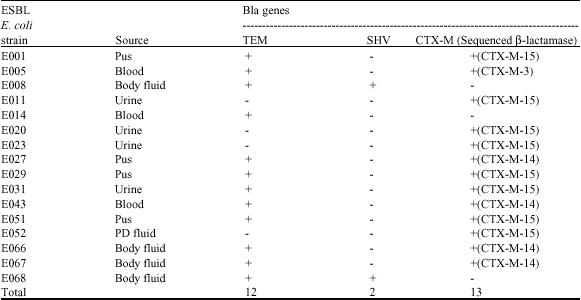
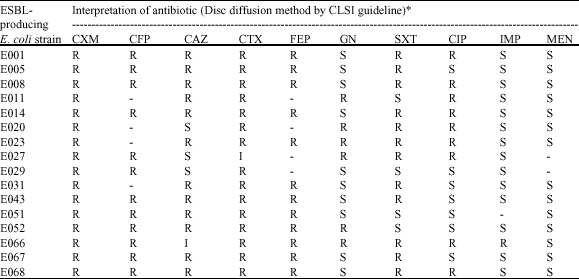
Extended-Spectrum β-Lactamases-Producing Escherichia colifrom a Tertiary Hospital in Malaysia: Emergence of CTX-M-Type β-Lactamases Variation
Department of Medical Microbiology and Parasitology,Faculty of Medicine and Health Sciences,Universiti Putra Malaysia, 43400 UPM Serdang, Selangor DE, Malaysia
Rusmah Yusof
Department of Medical Microbiology and Parasitology,Faculty of Medicine and Health Sciences,Universiti Putra Malaysia, 43400 UPM Serdang, Selangor DE, Malaysia
Mariana Nor Shamsudin
Department of Medical Microbiology and Parasitology,Faculty of Medicine and Health Sciences,Universiti Putra Malaysia, 43400 UPM Serdang, Selangor DE, Malaysia