ABSTRACT
Rhizobium species F1 was studied for its ability to grow in the presence of trehalose (Trehalose-Minimal Medium (TMM)) and absence of it (Nutrient Broth (NB)) as sole carbon and energy sources and form Trehalose-Catabolic-Enzyme (TCE). The organism was mutagenized with hydroxylamine. The resultant mutants and the parental strain were grown with and without trehalose. The supernatants of the grown culture alone and lysed cells in supernatants were assayed for the activity of TCE. Rhizobium species F1 and the mutants grew in TMM and NB. Many of the mutants grew better (OD670 = 0.36-1.0 in TMM and OD670 = 0.005-0.99 in NB) than the wild-type (OD670 = 0.51 in TMM and OD670 = 0.25 NB). All the strains constitutively and inducibly expressed the trehalose-catabolic enzyme with a range of 0.242-1.42 mg mL-1 glucose mg-1 mL-1 protein for the mutants and 1.025 mg mL-1 glucose mg-1 mL-1 protein for the parental type. In the absence of trehalose in the growth medium, the mutants synthesized higher amount of the TCE with the highest value of 1.091 mg mL-1 glucose mg-1 mL-1 protein and then the wild-type which exhibited enzyme activity of 0.321 mg mL-1 glucose mg-1 mL-1 protein. The enzyme was extracellularly and intracellularly expressed in the TMM and NB. Activity of the total trehalose-degrading enzyme was higher than that of the extracellular. Three classes of the mutants were identified. Low, normal and super-trehalose-catabolic-enzyme producers showed enzyme activity in the ranges of 0 to 30, 31 to 60 and above 60 mg mL-1 glucose mg-1 mL-1 protein, respectively.
PDF Abstract XML References Citation
How to cite this article
URL: https://scialert.net/abstract/?doi=jm.2008.269.275
INTRODUCTION
Trehalose is a non-reducing sugar made up of α-D-glucopyranosyl-α-D-glucopyranoside. It occurs in many living cells including some plants, fungi, actinomycetes, bacteria and some invertebrates (Elbein, 1974; Willis and Walker, 1999; Dardanelli et al., 2000). It is wide spread in symbotic organelles such as actinorhizal nodules, ectomycorrhiza and nitrogen-fixing nodules (Harley and Smith, 1983; Lopez and Torrey, 1985; Streeter, 1985).
Trehalose plays various roles in organisms. The sugar helps to preserve the structure of cell membrane and protein in the absence of water and the flavour, colour and texture of foods associated with fresh counterparts. In plants, it shows strong effect on growth and metabolism by inducing starch formation. It also plays a role in regulating carbohydrate allocation (Muller et al., 2001) and modulate sugar mediated gene expression (Wingler et al., 2000). Karim et al. (2007) showed that accumulating trehalose in tobacco produced stress tolerance.
Trehalose protects microbial cells. Martinez-Esparza et al. (2007) in a study with a tps1/tps1 trehalose-deficient mutant of Candida albicans, found out that the trehalose played a significant role in resistance to macrophage killing of the yeast. Louis et al. (1994) observed highest survival rates of Escherichia coli K12 and E. coli NISSLE 1917 in the presence of trehalose and sucrose among other compatible dissacharide solutes tested during drying and storage of the microorganisms. Intracellular trehalose and trehalose-metabolizing enzymes found in Candida albicans revealed that significant amounts of the disaccharide are stored in response to thermal and oxidative challenge of the organism (Pedreno et al., 2007). The yeast displayed higher resistance to environmental insults, such as heat shock (42°C) or saline exposure (0.5 M NaCl) and to both mild and severe oxidative stress (5 and 50 mM H2O2), which are relevant during in vivo infections. Wolf et al. (2003) showed that three pathways for trehalose metabolism in Cornebacteriuim glutamicum ATCC 13032 are significant in response to osmotic stress. Trehalose can be used in enzyme purification protocols to increase the purity and quality of the isolated protein and to perform structural studies (Sampedro et al., 2007).
Despite the positive roles played by trehalose, the sugar exhibits toxicity in certain plants (Veluthambi et al., 1981). It is important to detoxify the sugar by hydrolysis particularly in plant-microbe interactions. Acid hydrolysis of trehalose to two molecules of 2,3,4,6-tetra-O-methyl-D-glucose in non-living medium is possible (Finar, 1986). In living systems however, enzymes which metabolise trehalose are known. Phosphotransferase, amylotrehalase and similar enzymes which are specific for the synthesis, uptake and split of the dissacharide have been reported in Salmonella typhimurium (Postma et al., 1986), Bacillus popillae (Bhumiratana et al., 1974), E. coli (Boos et al., 1990) and rhizobia (Streeter, 1985). Sinorhizobium meliloti genomic library contains a DNA segment encoding enzyme which allows Ralstonia eutropha to grow on the α-glucosides sucrose, maltose and trehalose (Willis and Walker, 1999). In another symbiotic interaction, activities of synthetic and hydrolytic enzymes involved in trehalose metabolism were discovered in peanut rhizobial strains grown in control, hypersaline and non-ionic hyperosmotic media (Dardanelli et al., 2000). Cultured Bradyrhizobium japonicum and B. elkanii were found to have three enzymes for trehalose synthesis: Trehalose Synthase (TS), maltooligosyltrehalose synthase (MOTS) and trehalose-6-phosphate synthetase (Streeter and Gomez, 2006). According to them, the presence of three totally independent mechanisms for the synthesis of trehalose by Bradyrhizobium species suggests that this disaccharide is important in the function of this organism both in the free-living state and in symbiosis. Recently, I discussed the breakdown of trehalose by rhizobia and characterized the trehalose-degrading enzyme isolated from Rhizobium spp. NGR234, a broad-host rhizobium (Boboye, 2004). In rhizobia, certain studies have been documented on rhizobial trehalose-metabolic enzymes. In tropical rhizobia only few literatures are found on trehalose-degrading enzymes. Here, we reported that (1) a tropical Rhizobium species F1 synthesizes a trehalose-catabolic-enzyme and (2) hydroxylamine mutation affected the enzyme production. The findings obtained from this study could serve as a basis for further studies on the relevance of trehalose in tropical rhizobia-legume symbiosis since sugars are important in this relationship. This is synonymous to more researches that are being carried out on the most abundant carbon source (photosynthetically produced sucrose) transported into legume root nodules despite that the importance of its metabolism by rhizobia in planta is not yet known (Willis and Walker, 1999).
MATERIALS AND METHODS
Growth Media
Nutrient broth and agar were obtained from Lab M., Topley House, England. Trehalose Minimal Medium (TMM) contained 1.5% (w/w) peptone water and 2% (w/w) trehalose. The bacterium was grown at 28°C for 24 h and maintained on nutrient agar containing malachite green at 0.03% (w/v).
Mutation Experiment
Two milliliters of 15 h grown culture of the rhizobium (OD670 of 0.2) was mutated by the method of Parkinson (Parkinson, 1976) with slight modification. An aliquot (0.1 mL) of 0.5 M hydroxylamine was mixed with the culture, left at 28°C for about 1 h. Cells were spun down at 12.168x103 g for 15 min (MSE Minor 35 centrifuge). Cells were washed twice with rhizobial minimal medium (Broughton and Dilworth, 1971). They were resuspended in 20 mL nutrient broth, kept at 4°C for 18 h for the mutants to segregate. Aliquot of serially diluted cells suspension were inoculated into nutrient agar by standard pour plate technique. Killing and survival rates of the mutation were calculated relative to the colony counts of the wild-type strain. Master plate was made for the mutants and they were used for various tests.
Test for Effect of Trehalose on Growth
Growth in the absence and presence of trehalose was tested for by inoculating 5 mL of fresh nutrient broth and TMM separately with 10 μL of a 18 h grown culture of the microbe at OD670 of 0.48. Incubation was carried out at 28°C for 24 h. Optical density was read at 670 nm before and after incubation. This procedure was carried out using the mutants and the wild-type strains to examine the effect of mutation on the utilization of trehalose.
Preparation of Enzyme
Enzyme was obtained from the medium by centrifuging 1.0 mL of the grown culture at centrifugal force of 12168x103 g for 15 min (MSE Minor 35 Centrifuge). The supernatant was used as a source of extracellular enzyme. Cells were toluenized according to the method of Moran and Starr (1969) as described by Boboye and Shonukan (1993) with slight modification. Toluene (0.1 mL) was added to 4 mL TMM grown culture, shaken at 80 rpm for 25 min at 28°C in a water bath. It was spun as described earlier and the supernatant constituted the total enzyme source.
Enzyme Assay
This was carried out by adding enzyme source (0.5 mL), phosphate buffer at pH 7.0 (0.5 mL) and 2% (w/v) trehalose (0.2 mL) to distilled water (0.8 mL). This reaction mixture was incubated at 37°C for 1 h and boiled for 5 min to stop the reaction. Amount of glucose released by the enzyme was determined by using slightly modified standard method of determining dinitrosalicylic acid (DNSA). One milliliters of DNSA reagent was added, boiled for five minutes after which 9 mL of distilled water was added, cooled and the absorbance was read at 540 nm. Glucose standard curve was used to estimate the amount of glucose released during enzyme reaction in mg mL-1. Protein concentration in each enzyme preparation was estimated in milligram by standard Biuret method (Gornall et al., 1949). Trehalose-catabolic-enzyme activity was defined as the amount of glucose (mg mL-1) released during enzyme reaction mg-1 mL-1 protein.
RESULTS AND DISCUSSION
Rhizobium spp. F1 grew considerably in both nutrient broth and TMM but faster in the former with average better growth in the latter medium. At 24 h of incubation, the bacterium showed OD670 of 0.510 and 0.250 in the respective medium. Mutant strains of the organism showed similar pattern of growth in the media within the OD670 of 0.36 (MU 44, 46) to 1.0 (MU 38) in nutrient broth and 0.005 (MU 4) to 0.990 (MU 28) in TMM (Fig. 1). This indicates that F1 was encouraged to grow more in the nutrient broth than in the trehalose minimal medium because the former is a complete medium. The ability of the rhizobium and its mutants to grow in the presence of trehalose is an indication of a digestive enzyme/s synthesized which enabled them to use the sugar for metabolism in TMM.
Relative to the wild-type Rhizobium spp. F1, 0.5% colonies survived the chemical mutation. The high killing effect (99.5%) of the hydroxylamine is an indication that the mutation has caused some changes in the genome of the bacterium which could affect cellular mechanisms. The wild-type and mutant strains showed trehalose-catabolic-enzyme activity at varying levels in supernatants of nutrient broth and TMM grown cells (Fig. 2).
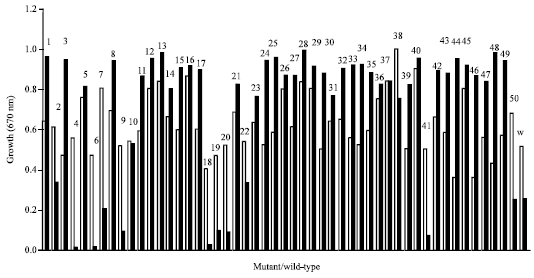 | |
| Fig. 1: | Effect of trehalose on the growth of mutant/wild-type strains of type of Rhizobium species F1. (□): growth in trehalose minimal medium and (■): growth in nutrient broth. Numbers 1-50 refer to the mutants and W is the wild-type strain of the Rhizobium species F1 |
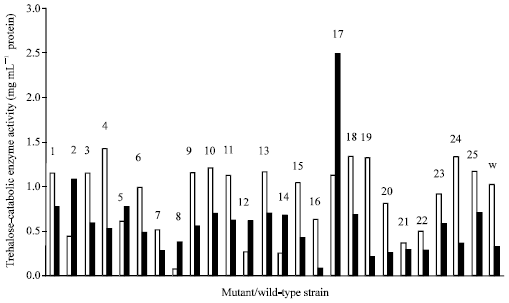 | |
| Fig. 2: | Trehalose-catabolic-enzyme (extracellular) activity of mutant/wild-type strains of Rhizobium species F1. (□): growth in trehalose minimal medium and (■): growth in nutrient broth. Numbers 1-25 refer to the mutants and W is the wild-type strain of the Rhizobium species F1 |
All the mutants with the exception of MU 2, 5, 8, 12 and 14 and the parental strain showed higher enzyme activity when grown in TMM than in nutrient broth with MU 4 expressing the highest activity of 1.42 mg mL-1 glucose mg-1 mL-1 protein in TMM and 0.529 mg mL-1 glucose mg-1 mL-1 protein in NB. The lowest enzyme activity exhibited by the mutants in TMM was 0.075 mg mL-1 glucose mg-1 mL-1 protein and 0.077 mg mL-1 glucose mg-1 mL-1 protein in NB. The trehalose breakdown activity of the wild-type was 1.025 mg mL-1 glucose mg-1 mL-1 protein in TMM and 0.321 mg mL-1 glucose mg-1 mL-1 protein in NB. This data (Fig. 2) implies that there was an enhancement of the synthesis of the enzyme by trehalose and not by the latter medium. It also shows that the enzyme was constitutively and extracellularly produced.
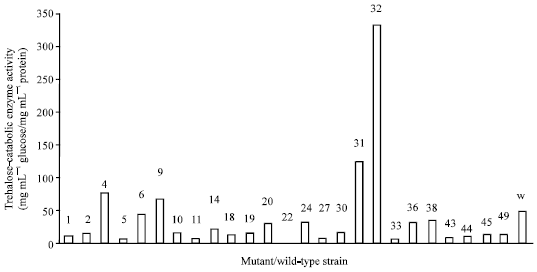 | |
| Fig. 3: | Total trehalose-catabolic-enzyme activity of mutant and wild-type strains of Rhizobium species F1. F1. Numbers 1-49 refer to the mutants and W is the Wild-type strain of the Rhizobium species F1 |
The other mutants (MU 2, 5, 8, 12, 14 and 17) which production of trehalose-catabolic-enzyme was not induced by trehalose appeared to have experienced mutational effect which block the synthesis at higher level. Total trehalose-catabolic enzyme activity was higher than the extracellular one (Fig. 3, 2). The difference in these activities indicates production of intracellular trehalose-catabolic-enzyme. Extracellularly expressed enzyme appeared lower than intracellular TCE in action; an indication that all the strains secreted a higher percentage of the enzyme within than outside the cell. The wild-type produced 97.9% of the total enzyme in its cytoplasm. The data obtained in this study is similar to that of some scientists. In Scytalidium thermophilum (a thermophilic fugus), trehalase was produced both extracellularly and intracellularly (Kadowski et al., 1995). Boboye (2004) reported that rhizobia synthesize trehalose-degrading enzyme. In her study, she showed that Rhizobium species NGR234, a temperate rhizobium did not produce trehalose-degrading enzyme extracellularly but intracellularly in contrast to this investigation. Also, she showed a result similar to this that the synthesis of the trehalose-degrading enzyme of NGR234 was induced by trehalose and not by succinate used in the experiment. Sinorhizobium meliloti Rm1021 synthesized ThuA and ThuB important for a major trehalose catabolism and are induced by trehalose but not by related structurally similar disaccharides like sucrose or maltose (Jensen et al., 2005). In this study, trehalose enhanced the formation of the enzyme. This is supported by the fact that enzymes involved in the breakdown of carbon and energy sources are often inducible. Enzyme induction is mediated by substrate or their analogs (Brock et al., 1984).
Based on the total trehalose-catabolic-enzyme activity of the mutants relative to that of the wild-type strain, the mutants were grouped into three classes namely: low, normal and super trehalose-catabolic-enzyme producers with activity of the enzyme ranging from 0 to 30, 31 to 60 and above 60 mg mL-1 glucose mg-1 mL-1 protein for the respective class. The normal producers formed the enzyme at a level similar to the wild-type (48.166 mg mL-1 glucose mg-1 mL-1 protein). The mutation did not considerably alter the genetic constitution of the enzyme-coding-gene in these normal producers. The higher and lower enzyme activities of some mutants than the wild-type was the resultant effect of the mutation on the gene’s encoding the synthesis of the enzyme or mutations in other genes which could indirectly affect the putative trehalose-degradation that was measured in this experiment. Hydroxylamine causes mutation by breaking DNA strand through indirect mechanism of facilitating depurination or depyrimidation during which cytosine undergoes a tautomeric shift pairing with adenine base (Fisbein et al., 1970).
Expression of genes including production of enzymes is regulated by induction and repression. Some genes are constitutively expressed at constant levels regardless of the cellular metabolic rate. In this experiment, the three regulatory processes occurred which means that trehalose induced the synthesis of the enzyme. Inducer (trehalose) attaches to repressor protein to deactivate it from repressing. Trehalose then sits on the operator locus of DNA causing the formation of mRNA which codes for the TCE (Brock et al., 1984). The wild-type, normal and super producers of the catabolic enzyme were induced at genetic and physiological levels. The hydroxylamine mutation caused alteration of the operator region of the gene such that the DNA did not respond to the action of the repressor protein thus, the normal producing mutants synthesized the enzyme within the range of that of the parental bacterium. The super producers appeared impaired in the genetic constitution of the repressor protein and did not produce the repressor anymore allowng the trehalose to freely induce synthesis of the enzyme above normal level of the wild-type (Madigan et al., 2001). The mechanism of breakage of the DNA strands by the hydroxylamine (Fisbein et al., 1970) could also make it possible to have more than normal number of the gene encoding TCE in the genome thus forming huge amount of the enzyme in the mutants as observed in the activity.
This research has shown that the tropical Rhizobium species F1 and its mutants formed trehalose-catabolic-enzyme when cultured in the presence and absence of trehalose and supernatants of the growth media and within the cytoplasm. Chemical mutation of the bacterium yielded low, wild-type-like and super producers of trehalose-catabolic-enzyme. Sequence analysis of DNA of representative members of the mutant classes and parental strain should be investigated to reveal the order of nucleotides thereby locating the real loci of the mutation.
ACKNOWLEDGMENTS
We thank the Department of Microbiology and Mr. Oseni in the Department of Biochemistry of the Federal University of Technology, Akure, Ondo State, Nigeria for instrumentation and chemicals used for this study.
REFERENCES
- Boboye, B., 2004. Degradation of trehalose by rhizobia and characteristics of a trehalose-degrading enzyme isolated from Rhizobium species NGR234. J. Applied Microbiol., 97: 256-261.
Direct Link - Dardanelli, M.S., P.S. Gonzalez, M.A. Bueno and N.E. Ghittoni, 2000. Synthesis, accumulation and hydrolysis of trehalose during growth of peanut rhizobia in hyperosmotic media. J. Basic Microbiol., 40: 149-456.
Direct Link - Jensen, J.B., T.V. Bhuvaneswari, O.Y. Ampomah, R. Darrah and N.K. Peters, 2005. Role of trehalose transport and utilization in Sinorhizobium meliloti-alfalfa interactions. Mol. Plant Microbe Interact., 18: 694-702.
Direct Link - Kadowaki, M.K., M. de Lourdes, T.M. Polizeli, H.F. Terenzi and J.A. Jorge, 1995. Characterization of trehalase activities from the thermophilic fungus Scytalidium thermophilum. Biochem. Biophy. Acta (BBA)-Gen. Subj., 1291: 199-205.
CrossRefDirect Link - Karim, S., H. Aronsson, H. Fricson, M. Pirthonen and B. Leyman et al., 2007. Improved drought tolerance without undesired side effects in transgenic plants producing trehalose. Plant Mol. Biol., 64: 371-386.
Direct Link - Lopez, M.F. and J.G. Torrey, 1985. Purification and properties of trehalase in Frankia ArI3. Arch. Microbiol., 143: 209-215.
CrossRefDirect Link - Louis, P., H.P. Truper and E.A. Galinski, 1994. Survival of Escherichia coli during drying and storage in the presence of compatible solutes. Applied Microbiol. Biotechnol., 41: 684-688.
CrossRefDirect Link - Martinez-Esparza, M., A. Aguinaga, P. Gonzalez-parraga, P. Garcia-penarrubia, T. Jouault and J.C. Arguelles, 2007. Role of trehalose in resistance to macrophage killing: Study with a tps1/tps1 trehalose-deficient mutant of Candida albicans. Clin. Microbiol. Infect., 13: 384-394.
Direct Link - Moran, F. and M.P. Starr, 1969. Metabolic regulation of polygalacturonic acid transeliminase in Erwinia. Eur. J. Biochem., 11: 291-295.
CrossRefDirect Link - Muller, J., R.A. Aeschbacher, A. Wingler, T. Boller and A. Wiemken, 2001. Trehalose and trehalase in Arabidopsis. Plant Physiol., 125: 1086-1093.
Direct Link - Parkinson, J.S., 1976. CheA, CheB and CheC genes of Escherichia coli and their role in chemotaxis. J. Bacteriol., 126: 758-770.
Direct Link - Pedreno, Y., P. Gonzalez-parraga, M. Martinez-esparza, R. Sentandreu, E. Valentin and J.C. Arguelles, 2007. Disruption of the Candida albicans ATC1 gene encoding a cell-linked acid trehalase decreases hypha formation and infectivity without affecting resistance to oxidative stress, Microbiology, 153: 1372-1381.
Direct Link - Postma, P.W., H.G. Keizer and P. Koolwijk, 1986. Transport of trehalose in Salmonella typhimurium. J. Bacteriol., 168: 1107-1111.
Direct Link - Streeter, J.G. and M.L. Gomez, 2006. Three enzymes for trehalose synthesis in Bradyrhizobium cultured bacteria and in bacteroids from soybean nodules. Applied Environ. Microbiol., 72: 4250-4255.
Direct Link - Wingler, A., T. Fritzius, A. Wiemken, T. Boller and R.A. Aeschbacher, 2000. Trehalose induces the ADP-glucose pyrophosphorylase gene, APL3 and starch synthesis in Arabidopsis. Plant Physiol., 124: 105-114.
CrossRefDirect Link - Wolf, A., R. Krämer and S. Morbach, 2003. Three pathways for trehalose metabolism in Cornebacteriuim glutamicum ATCC 13032 and their significance in response to osmotic stress. Mol. Microbiol., 49: 1119-1134.
Direct Link








