Research Article
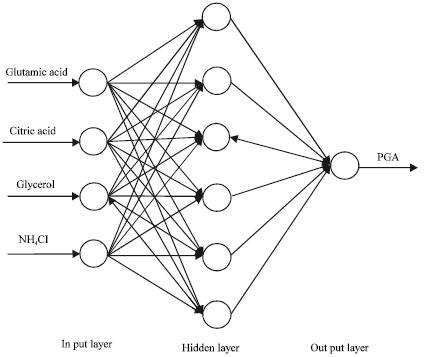
Optimization of Culture Medium for the Production of Poly-γ-glutamic Acid Using Artificial Neural Networks and Genetic Algorithms
Department of Chemical Engineering, Center for Biotechnology, Andhra University, Visakhapatnam-530 003, India
Garapati Hanumantha Rao
Department of Chemical Engineering, Center for Biotechnology, Andhra University, Visakhapatnam-530 003, India