Research Article
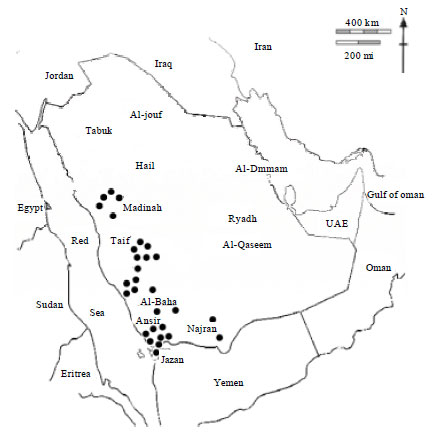
Population Structure of the Yemeni Honey Bee (Apis mellifera jemenitica) Entails an Urgent Conservation Strategy in Saudi Arabia
Baqshan Chair for Bee Research, Faculty of Food Science and Agriculture, King Saud University, Riyadh, 11451, Saudi Arabia
A. AlGhamdi
Baqshan Chair for Bee Research, Faculty of Food Science and Agriculture, King Saud University, Riyadh, 11451, Saudi Arabia
M. Alsharhi
Baqshan Chair for Bee Research, Faculty of Food Science and Agriculture, King Saud University, Riyadh, 11451, Saudi Arabia