Research Article
Genetic Biodiversity, Affinities and Phylogeny of Four Goat Breeds in Egypt
Department of Cell Biology, National Research Centre, Dokki, Egypt
Eman R. Mahfouz
Department of Cell Biology, National Research Centre, Dokki, Egypt
Domestic goats (Capra hircus) are economically important livestock especially in developing countries like Egypt. Goat is considered to be one of the important sources of milk, meat, hair and fiber (Porter, 1996). On the other hand, goat can tolerate the harsh environmental conditions like drought and poor nutrition (Joshi et al., 2004).
Some archaeological studies showed that the place of goat domestication is the Fertile Crescent region 10,000 years ago (Pringle, 1998; Zeder and Hesse, 2000). Another theory suggested that the domestication of goat originated from Pakistan and derived from Cashmere goat breeds (Meaow, 1993). Till now, the origins of domestic goats are still not well documented in spite of its crucial and important role for human beings since very old history around Neolithic times (Legge, 1996; Porter, 1996, Pringle, 1998).
Mitochondrial DNA (mtDNA) is considered to be the most informative genomic data for declaring the origins of domestic livestock (MacHugh and Bradley, 2001). Till now, mitochondrial sequences have been widely used to study the origin of cattle (Loftus et al., 1994; Bradley et al., 1996; Troy et al., 2001), sheep (Hiendleder et al., 1998, 2002), goat (Luikart et al., 2001; Mannen et al., 2001; Sultana et al., 2003; Joshi et al., 2004; Doro et al., 2014), pig (Giuffra et al., 2000), dog (Vila et al., 1997; Savolainen et al., 2002), horse (Vila et al., 2001) and donkey (Beja-Pereira et al., 2004).
Previous reports mentioned that there are 4 major mtDNA lineages of domestic goats. Lineage A is the most common and is distributed all over the world. Lineage B is restricted to Southern and Eastern Asia, including Pakistan, India, Mongolia and Malaysia. Lineage C is found in low frequencies in some countries like Switzerland, Slovenia, Pakistan and India. Finally, lineage D is very rare and is observed only in some regions of Pakistan and India (Luikart et al., 2001; Sultana et al., 2003; Joshi et al., 2004). Naderi et al. (2007) reported the presence of two other haplogroups: F and G, which are present with few frequencies and in restricted localizations.
Using the sequences of mtDNA control region, this work aimed to analyze and clarify the genetic biodiversity and affinities of four goat breeds reared in Egypt. This study could also indicate the plylogeny of these breeds and therefore it contributes in understanding the goat domestication history.
Blood samples and genomic DNA extraction: Blood samples were collected, in a period of time ranging from January 2014 to July, 2015, from goat animals belonging to 4 main goat breeds reared in Egypt namely; Barki, Baladi, Zaraiby and Damascus. Genomic DNA was extracted from the whole blood according to the method described by Miller et al. (1988) with minor modifications. Briefly, Blood samples were mixed with cold 2x sucrose-triton and centrifuged at 5000 rpm for 15 min at 4°C. The nuclear pellet was suspended in lysis buffer, sodium dodecyl sulfate and proteinase K and incubated overnight in a shaking water bath at 37°C. Nucleic acids were extracted with saturated NaCl solution. The DNA was picked up and washed in 70% ethanol. The DNA was dissolved in 1xTE buffer. The DNA concentration was determined, using Nano Drop1000 thermo scientific spectrophotometer and then diluted to the working concentration of 50 ng μL–1.
Polymerase Chain Reaction (PCR): The hypervariable segment (HVSI) of the mtDNA control region was amplified and sequenced. Primers were used to amplify a 579-bp DNA fragment (Luikart et al., 2001).
| CAP-F: | CGT GTA TGC AAG TAC ATA C |
| CAP-R: | CTG ATT AGT CAT TAG TCC ATC |
The PCR amplifications were conducted in a 50 μL volume containing 5 μL of 10x reaction buffer, 1.5 mM MgCl2, 0.2 mM dNTPs, 0.2 μM each primer, 1.5U Taq DNA polymerase and approximately 30 ng genomic DNA. The PCR mixture underwent 3 min at 95°C followed by 35 cycles of: 50 sec at 94°C, 1 min at 55°C, 1 min at 72°C and 5 min at 72°C. The PCR products were purified and sequenced.
Data analysis: The D-loop sequences were aligned using the BioEdit software (Hall, 1999) in order to identify and trace individual haplotype mutations.
The DnaSP 5.00 software (Librado and Rozas, 2009) was used to calculate haplotype structure and DNA divergence between groups of haplotypes.
Neighbor-joining haplotype tree was computed with Mega version 5.0 (Tamura et al., 2011).
Goats are spread widely in many countries throughout the world and this livestock species possesses a high capability for adaptation in harsh environmental conditions including high temperature, drought and poor nutrient sources. These advantages in addition to disease tolerance make the goat one of the highly valuable economically important livestock in the world. Its biological features include an efficient reproductive system and small body sizes which enable it to survive in marginal environments. Goat are easy for management because they do not require high input and this advantage makes them the most suitable resource, in poor rural households, for multiple role including meat, milk, hair and fiber production (Acharaya and Battasha, 1992).
Threats to biodiversity are increasing due to the loss of genetic diversity within the species utilized in agriculture. The formulation of the modern breed concept has caused remarkable changes in the livestock sector and its application to breeding, while husbandry practices led to the formation of well-defined breeds and subsequently intense anthropogenic selection (Porter, 2002). Therefore, farmers progressively substituted the less productive, locally adapted native breeds with highly productive cosmopolitan ones and progressively abandoned marginal areas (Taberlet et al., 2008). Accordingly, it is more strategically important than ever to preserve as much farm animal diversity as possible, to ensure a prompt and proper response to the needs of future generations.
Among many livestock species, the variations in mtDNA control region has been the best tool to declare the biodiversity of present breeds and also to reconstruct the domestication time and places (Wang et al., 2008; Wu et al., 2009; Pereira et al., 2009; Vacca et al., 2010). This region of the genome possesses three advantages which make it provide excellent and optimal markers for diversity and phylogenic studies. It possesses sufficient evolutionary conservation, diversity, variability and structuring among the species and fast constant evolutionary rate (Bruford et al., 2003). In addition, the mtDNA contains a high number of copies per cell and this advantage allows possible analyses even from minute amounts of material and also its haploid manner makes genotyping clear without confusion with the highly polymorphic hypervariable (HV) segment (Piras et al., 2012).
In this study, 520-bp fragments from goat mtDNA control region were amplified using specific primers. These amplified fragments were sequenced and the nucleotide sequences of 70 samples belonging to 4 goat breeds were aligned using BioEdit software. The DnaSP 5.00 software was used to identify the sequence variation and polymorphic sites in the aligned sequences.
The results showed the presence of 69 polymorphic sites leading to the construction of 44 different haplotypes (Fig. 1). The sequences of all detected haplotypes in this study were submitted to GenBank database with the accession numbers KU145776-KU145819. Haplotypes No. 1 and 23 were the most detected ones and were found in five samples. Haplotype No.1 was shared in Barki and Baladi breeds, whereas, the five samples displaying haplotype No. 23 belonged to Damascus breed. Haplotype No. 33 appeared in 4 samples, all of which belonged to Zaraibi breed. Haplotypes No. 7, shared in Barki and Baladi breeds and No. 31, specific for Zaraibi breed, appeared in 3 samples. All other 39 haplotypes appeared in 1 or 2 samples.
Damascus breed displayed 7 specific haplotypes, not shared with any other breed, whereas Zaraibi breed showed 15 haplotypes and shared one of them with Barki breed. Barki breed displayed 8 specific haplotypes, seven of which were shared with Baladi breed, in addition to a single haplotype shared with Zaraibi breed. Baladi breed showed 6 haplotypes and shared 6 of them with Barki breed, which is the highest number of shared haplotypes.
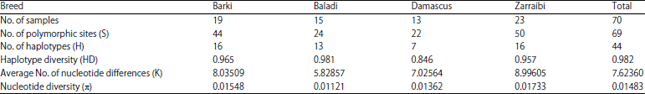
The analysis of sequence data of the 4 tested goat breeds (Table 1), using DnaSP 5.00, showed that Zaraibi and Barki breeds had the highest number of haplotypes (16 haplotypes) with 50 and 44 polymorphic sites, respectively. Baladi breed had 13 haplotypes with 24 polymorphic sites, whereas, Damascus breed possessed 7 haplotypes with 22 polymorphic sites.
The haplotype diversity in the 4 tested goat breeds ranged from 0.846 in Damascus breed (with an average number of nucleotide differences K: 7.02564) to 0.981 in Baladi breed (K: 5.82857). On the other hand, the haplotype diversity and average number of nucleotide differences in Zaraibi breeds were 0.957 and 8.99605, respectively, while in Barki breed were 0.965 and 8.03509, respectively. The results showed that the nucleotide diversity ranged from 0.01121 in Baladi breed to 0.01733 in Zaraibi breed with a total nucleotide diversity of 0.01483 in the 4 tested breeds.
The nucleotide difference (D) and the average number of pairwise differences (Dxy) between the tested breeds were estimated. The lowest D and Dxy values were observed between Baladi and Damascus (D: 6.717 and Dxy: 0.01302) followed by the differences between Barki and Damascus (D: 7.823 and Dxy: 0.01519), while the highest differences were observed between Damascus and Zaraibi (D: 8.521 and Dxy: 0.01654) followed by the differences between Barki and Zaraibi (D: 8.317 and Dxy: 0.01606). The D and Dxy values for Baladi and Zaraibi (D: 7.828 and Dxy: 0.01508) were found to be close to those of Barki and Damascus (Table 2).
The net distances between breeds were calculated using Mega 5.0 software. The results showed that the highest distances were recorded between Damascus and the other three breeds; Baladi, Barki and Zaraibi, descendingly (Table 3). This result is logically supported by the fact that Damascus breed is not considered as a local Egyptian breed, while it came from Syria, where it originated (Mavrogenis et al., 2006).
 | |
| Fig. 1: | Sequences of 44 haplotype sequences, the variable site numbers in red color |
| Table 1: | Genetic diversity data |
 | |
| Table 2: | Average pairwise differences between breeds |
 | |
| Average number of nucleotide difference between breeds D (below), average number of nucleotide substitution per site between breeds Dxy (above) | |
On the other hand, the net distances between Barki, Baladi and Zaraibi were small, which declares the presence of cross breeding between these three local Egyptian breeds.
| Table 3: | Net distance between breeds |
 | |
Neighbor-joining (phylogeny) tree was constructed using the Mega 5.0 software (Fig. 2). The sequences of the 70 tested samples were aligned with sequences of different reference haplogroups to detect the affinities of goat breeds in Egypt.
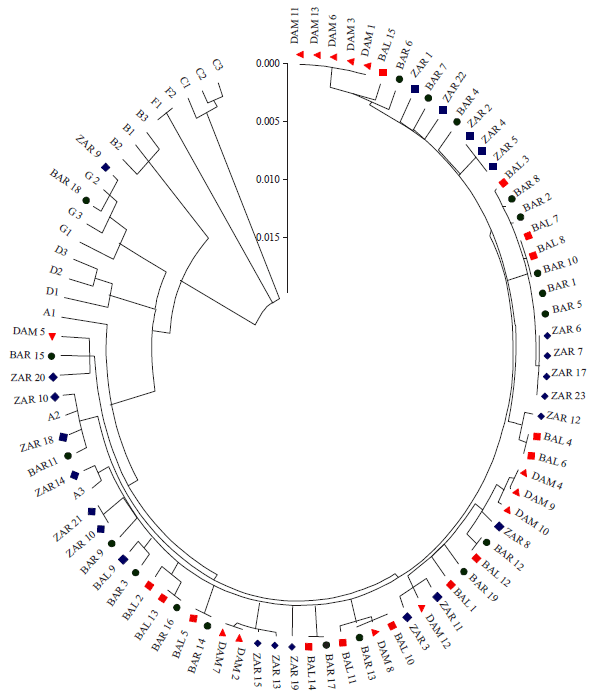 | |
| Fig. 2: | Neighbor-joining (NJ) tree of the tested animals as circle |
The reference sequences used were: EF617863 (A1), EF618309 (A2) and EF618492 (A3) for haplogroup A; EF618245 (B1), EF617706 (B2) and EF617850 (B3) for haplogroup B; EF617786 (C1), EF618486 (C2) and HQ596552 (C3) for haplogroup C; EF617701 (D1), EF618219 (D2) and AY155952 (D3) for haplogroup D; DQ241349 (F1) and DQ241351 (F2) for haplogroup F, while EF618084 (G1), EF618345 (G2) and EF618539 (G3) for haplogroup G.
The phylogeny results showed that all tested animals belong to haplogroup A with the exception of two samples; one from Barki and the other from Zaraibi breeds, which were found to belong to haplogroup G. This result agrees with Naderi et al. (2007), who reported the presence of six goat haplogroups with high diversity, where the examined 29 Egyptian goats were classified as 27 belonging to haplogroup A, while the remaining two belonged to haplogroup G.
The sequence references used in this study for haplogroup A were A1 (Iran), A2 (Saudi Arabia) and A3 (Turkey). Most of Egyptian goat animals tested belonged to haplogroup A, more specifically A3 and this observation confirms the hypothesis suggesting that the place of goat domestication is the Fertile Crescent region (Pringle, 1998; Zeder and Hesse, 2000), while the sheep interred North Africa via., Sinai. Also, Doro et al. (2014) sequenced the entire mtDNA of 28 Sardinian goats and found that all tested animals belonging to haplogoup A which is most common and is distributed all over the world.
In conclusion, the mtDNA in still the most informative tool for identification of biodiversity, affinities and phylogeny of different livestock breeds. Most Egyptian goats belong to haplogroup A, which is spread in many countries around the world while few samples belong to haplogroup G and are present in restricted areas including Iran and Turkey. The genetic distance is wider between Damascus breed and the other local Egyptian three breeds, where, the distance between them is narrow. These findings shed some light on the domestication place of goat which is suggested to be the Fertile Crescent region.