Research Article
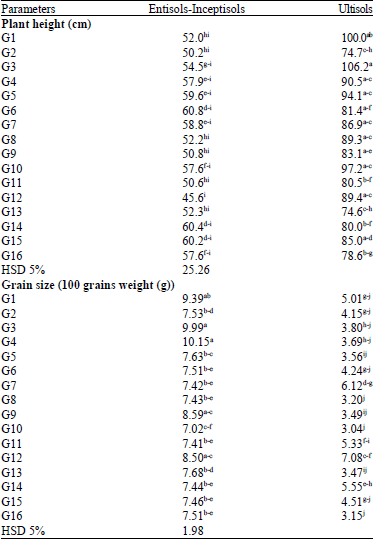
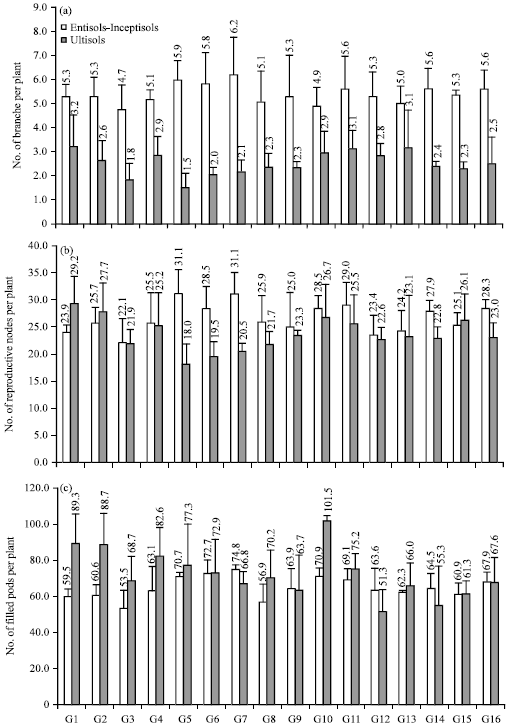
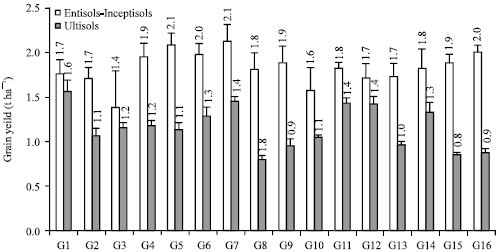
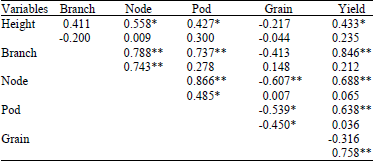
Decreasing Grain Size Caused Declining Grain Yield of CpMMV-Resistant Soybean Lines Grown in Ultisols
Indonesian Legume and Tuber Crops Research Institute, Indonesian Agency for Agricultural Research and Development, Jl. Raya Kendalpayak Km. 8, P.O. Box 66, Malang, 65101, Indonesia
Siti Zubaidah
Department of Biology, Faculty of Mathematics and Natural Sciences, Malang State University, Jl. Surabaya 6 Malang, 65145, Indonesia
Sulisetijono
Department of Biology, Faculty of Mathematics and Natural Sciences, Malang State University, Jl. Surabaya 6 Malang, 65145, Indonesia