Short Communication
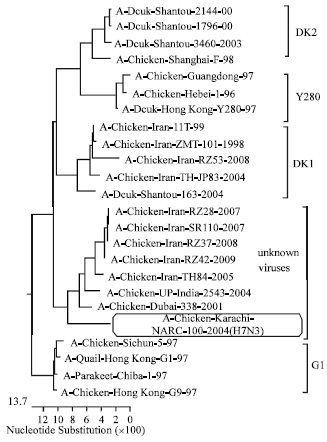
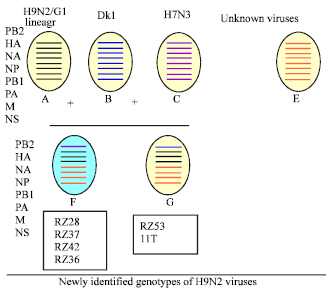
Characterization of PB2 Gene of H9N2 Avian Influenza Viruses from Iran, 2008 to 2009
Division of Avian Diseases,Islamic Azad University, Shoushtar Branch, Khozestan, Iran
Ali Bagherpour
Division of Microbiology, Islamic Azad University, Shoushtar Branch, Khozestan, Iran