Research Article
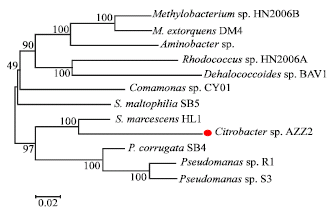
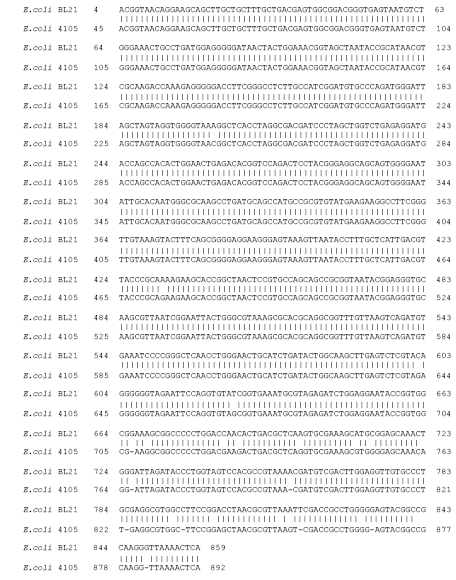
Molecular Prediction of Dehalogenase Producing Microorganism using 16S rDNA Analysis of 2,2-dichloropropionate (Dalapon) Degrading Bacterium Isolated from Volcanic Soil
Department of Industrial Biotechnology, Faculty of Biosciences and Bioengineering, Universiti Technologi Malaysia, 81310 Skudai, Johor, Malaysia
S. Hamdan
Faculty of Science and Technology, School of Biosciences and Biotechnology, Universiti Kebangsaan Malaysia, Selangor, Malaysia
S.H.Z. Ariffin
Faculty of Science and Technology, School of Biosciences and Biotechnology, Universiti Kebangsaan Malaysia, Selangor, Malaysia
F. Huyop
Department of Industrial Biotechnology, Faculty of Biosciences and Bioengineering, Universiti Technologi Malaysia, 81310 Skudai, Johor, Malaysia