Research Article
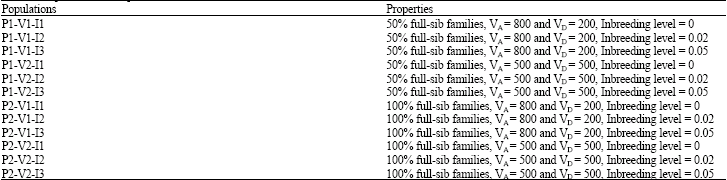
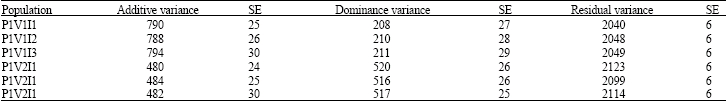
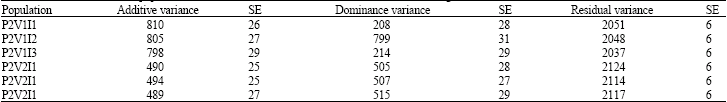
Estimates of Additive and Non-Additive Genetic Variances with Varying Levels of Inbreeding
Department of Animal production, School of Agricultural and Environmental Sciences, University of Limpopo, P/Bag X1106, Sovenga, South Africa
W. Ngambi
Department of Animal production, School of Agricultural and Environmental Sciences, University of Limpopo, P/Bag X1106, Sovenga, South Africa
C.A. Mbajiorgu
Department of Animal production, School of Agricultural and Environmental Sciences, University of Limpopo, P/Bag X1106, Sovenga, South Africa