Research Article
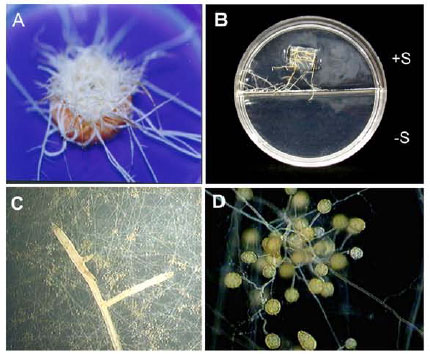
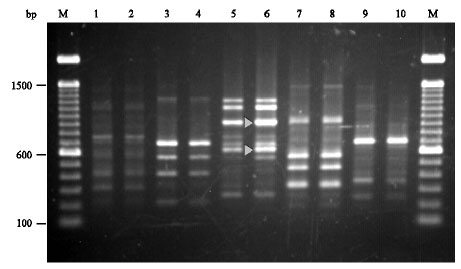
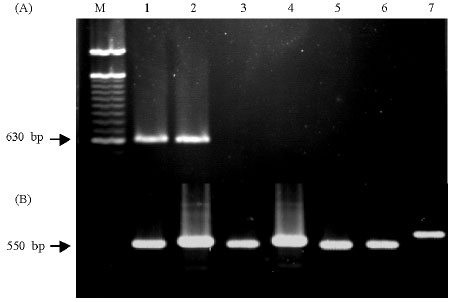
Specific Polymerase Chain Reaction-Based Assay for the Identification of the Arbuscular Mycorrhizal Fungus Glomus intraradices
Laboratorio de Biologia Molecular, Instituto Tecnologico de Tlajomulco, Jalisco. Tlajomulco de Zuniga, Jalisco, Mexico
J. Lara-Reyna
Colegio de Postgraduados Campus Campeche, Campeche, Campeche Mexico
L.V. Hernandez-Cuevas
CICB, Universidad Autonoma de Tlaxcala, Tlaxcala, Tlaxcala, Mexico
J.P. Martinez-Soriano
CINVESTAV, Campus Guanajuato, Irapuato, Guanajuato, Mexico