Research Article
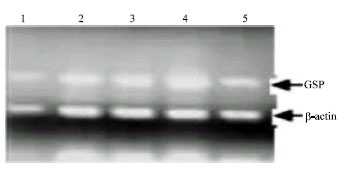
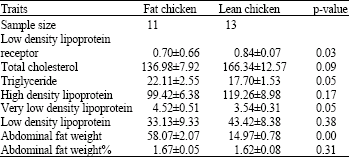
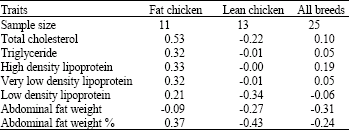
Analysis of LDL Receptor mRNA Expression, Serum Biochemical and Abdominal Fat Weight in Fat and Lean Chickens
College of Animal Science and Technology, Yangzhou University, Yangzhou, 225009, China
J.H. Cheng
College of Animal Science and Technology, Yangzhou University, Yangzhou, 225009, China
X.S. Wu
College of Animal Science and Technology, Yangzhou University, Yangzhou, 225009, China
H.P. Ju
College of Animal Science and Technology, Yangzhou University, Yangzhou, 225009, China
D.M. Mekki
College of Animal Science and Technology, Yangzhou University, Yangzhou, 225009, China
G.H. Chen
College of Animal Science and Technology, Yangzhou University, Yangzhou, 225009, China