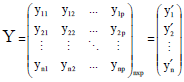Research Article
Outlier Detection for Multivariate Multiple Regression in Y-direction
School of Applied Statistics, National Institute of Development Administration,Bangkapi, Bangkok, 10240, Thailand
Pachitjanut Siripanich
School of Applied Statistics, National Institute of Development Administration,Bangkapi, Bangkok, 10240, Thailand