Research Article
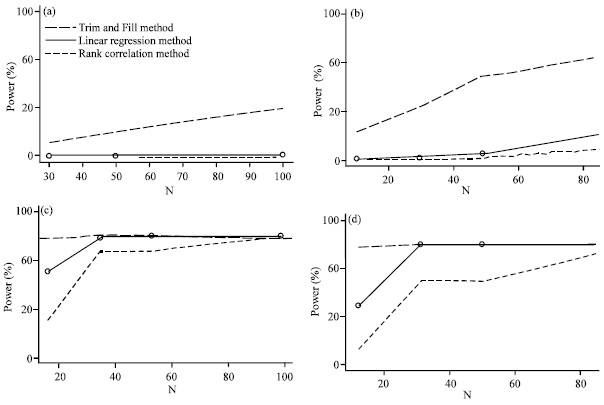
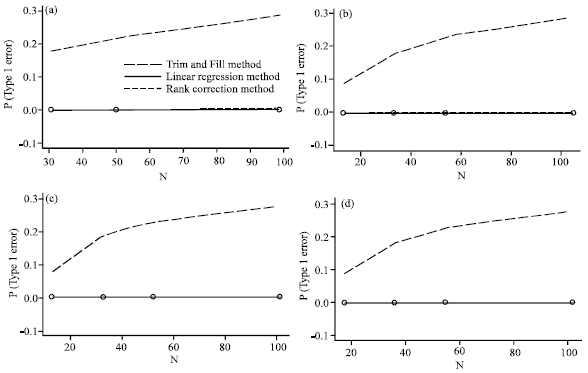
A Comparison of Methods to Detect Publication Bias for Meta-analysis of Continuous Data
Department of Computational and Theoretical Sciences, Kulliyyah of Science, International Islamic University Malaysia, Kuantan 25200, Pahang, Malaysia