Research Article
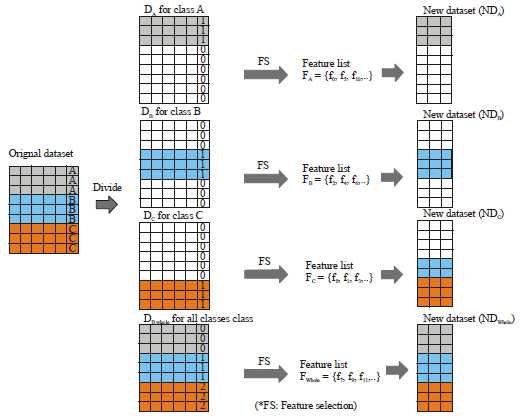
Divide and Merge Classification for High Dimensional Multi-Class Datasets
Department of Nanobiomedical Science, Dankook University, Cheonan, 330-714, South Korea
Sangbum Lee
Department of Computer Science, Dankook University, Cheonan, 330-714, South Korea