Research Article
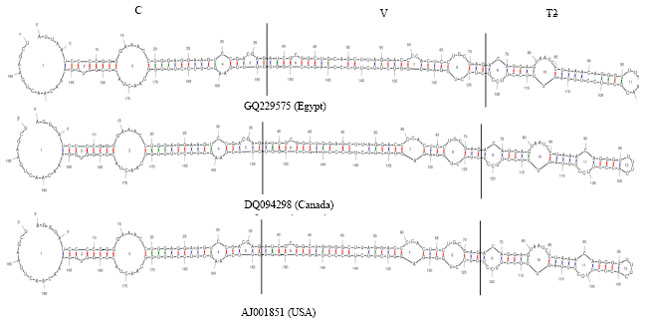
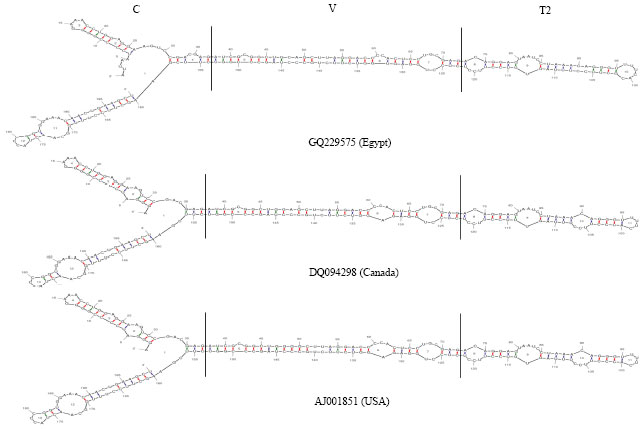
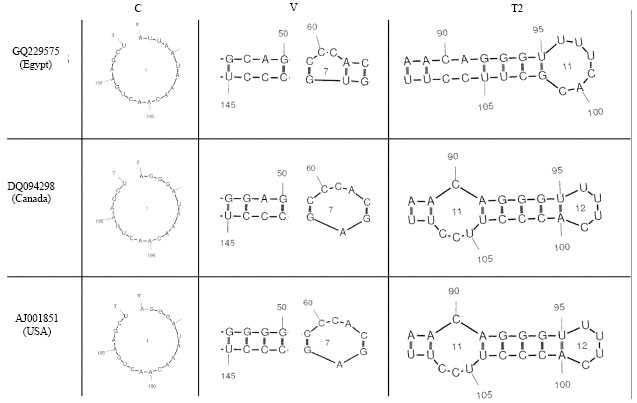
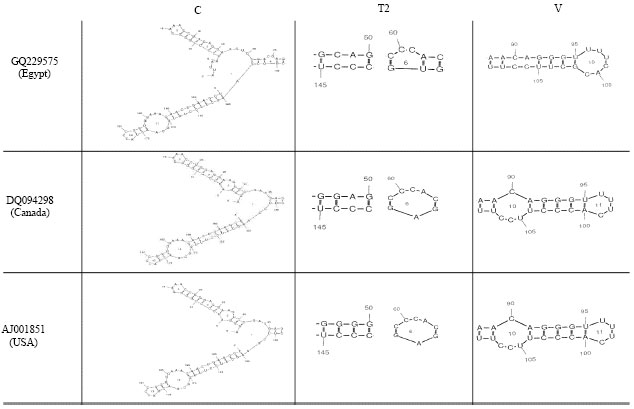
Partially Nucleotide Sequence and Secondary Structure of Chrysanthemum stunt viroid Egyptian Isolate from Infected-Chrysanthemum Plants
Virology Laboratory, Department of Agriculture Microbiology, Faculty of Agriculture, Ain Shams University, 11241 Cairo, Egypt
A.A. Rezk
Plant Virus and Phytoplasma Research Section, Plant Pathology Institute, Agriculture Research Center, Saudi Arabia
Dawoud A. Rehab
Plant Virus and Phytoplasma Research Section, Plant Pathology Institute, Agriculture Research Center, Saudi Arabia
A.R. Sofy
Department of Botany and Microbiology, Faculty of Science, Al-Azhar University, 11884 Nasr City, Cairo, Egypt