Research Article
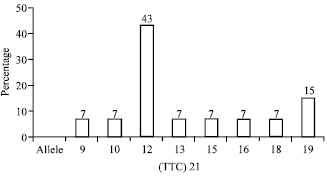
Strain Typing and Strain Differentiation of Mycobacterium leprae by TTC Repeats
School of Life and Health Sciences, Adikavi Nannaya University,Rajahmundry, 53510, Andhra Pradesh, India
Kalyani Dasari
School of Life and Health Sciences, Adikavi Nannaya University,Rajahmundry, 53510, Andhra Pradesh, India
Idress Hamad Attitalla
Department of Microbiology, Faculty of Science, Omar Al-Mukhtar University,P.O. Box 919, Al-Bayda, Libya
Srikantam Aparna
Blue Peter Research Centre, LEPRA Society, Cherlapally, Hyderabad, 501301, Andhra Pradesh, India
Padmaja Ponnada
School of Life and Health Sciences, Adikavi Nannaya University,Rajahmundry, 53510, Andhra Pradesh, India
Madan Mohan Male
School of Life and Health Sciences, Adikavi Nannaya University,Rajahmundry, 53510, Andhra Pradesh, India