Research Article
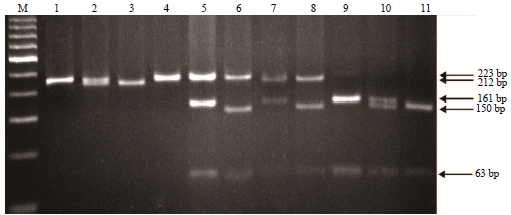
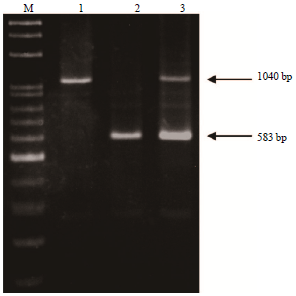
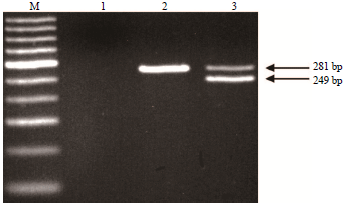
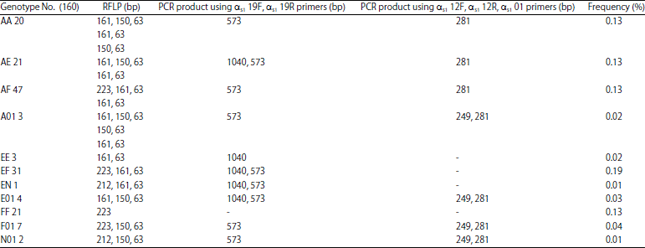
Genotyping of the CSN1S1 Locus in Taiwan Dairy Goats using PCR-RFLP and AS-PCR
Department of Animal Science, National Chiayi University, Chiayi, Taiwan
Shu- Yu Hsieh
Department of Animal Science, National Chiayi University, Chiayi, Taiwan
Yan- Ming Horng
Department of Animal Science, National Chiayi University, Chiayi, Taiwan
Chean- Ping Wu
Department of Animal Science, National Chiayi University, Chiayi, Taiwan