Research Article
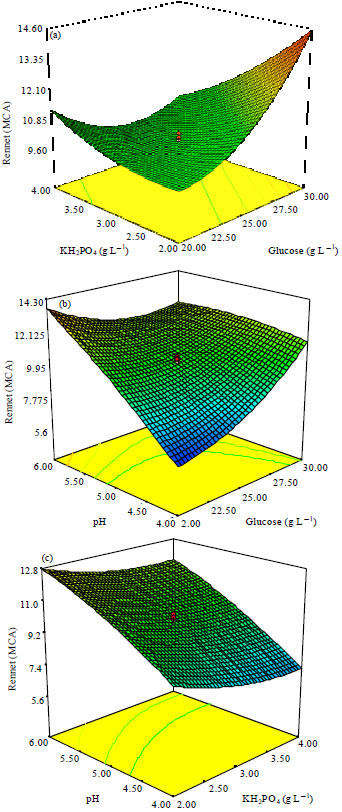
Statistical Optimization of Milk Clotting Enzyme Biosynthesis by Mucor mucedo Kp736529 and its Further Application in Cheese Production
Department of Dairy Research, Food Technology Research Institute, Agriculture Research Centre, Giza, Egypt
Amal Elsady Ibrahim
Department of Dairy Research, Food Technology Research Institute, Agriculture Research Centre, Giza, Egypt
Wesam I.A. Saber
Microbial Activity Unit, Department of Microbiology, Soils, Water and Environment Research Institute, Agricultural Research Center, P.N. 12619, Giza, Egypt