Research Article
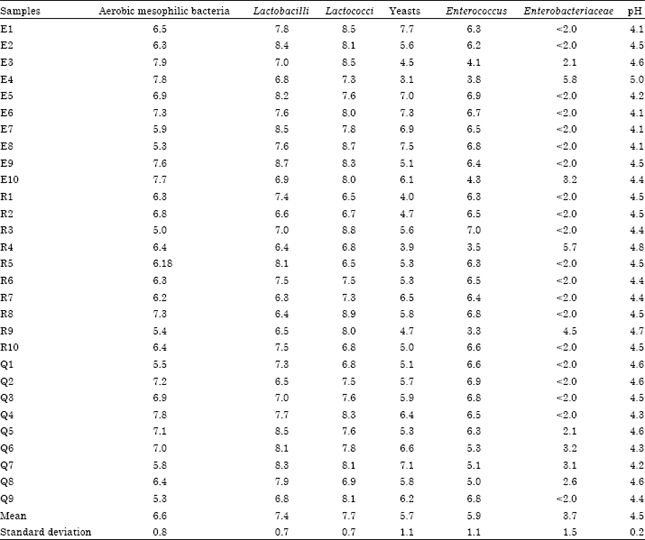
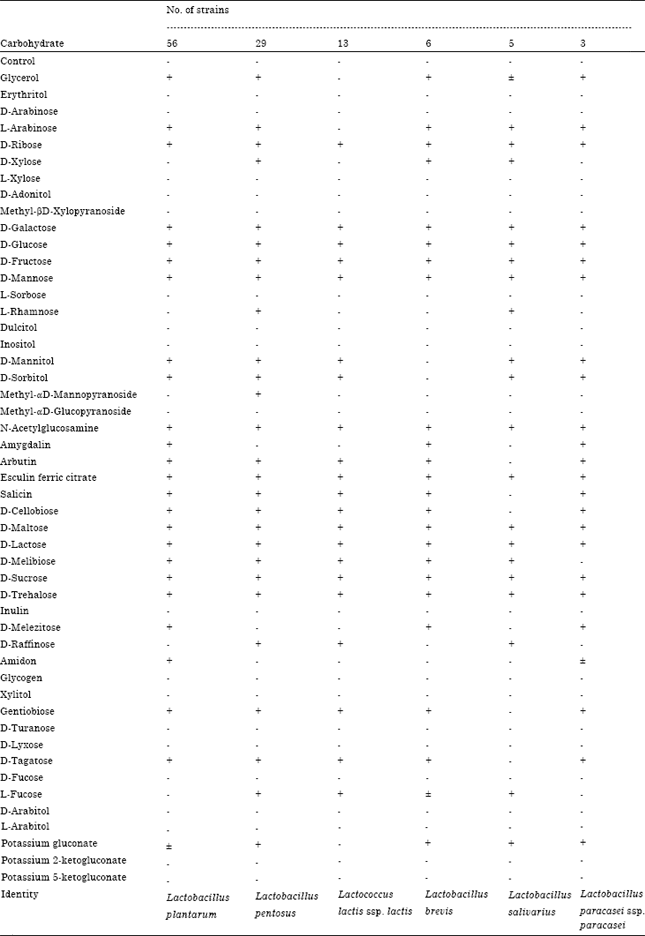
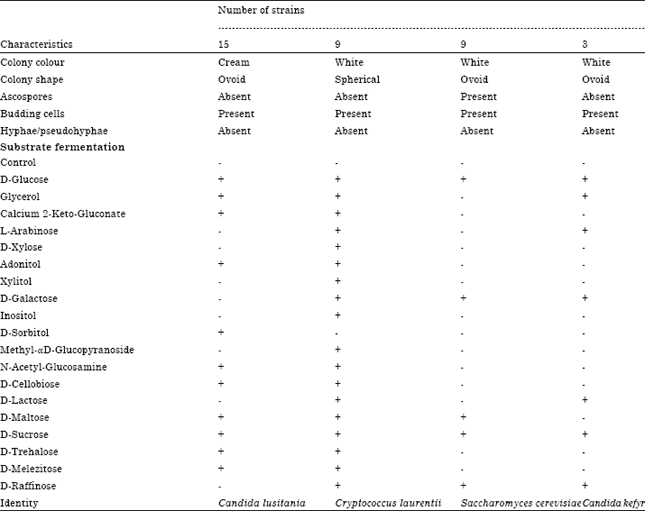
Isolation and Identification of Lactic Acid Bacteria and Yeasts from Sameel Milk: A Saudi Traditional Fermented Milk
Department of Food and Nutrition Sciences, College of Agriculture and Food Science, King Faisal University, P.O. Box 420 Al-Ahsa 31982, Saudi Arabia