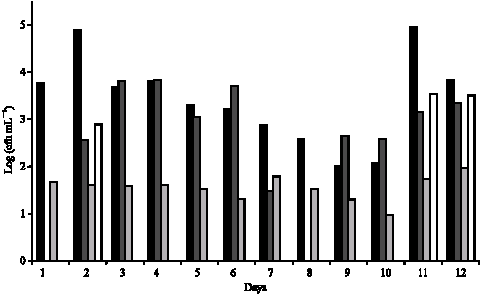Research Article
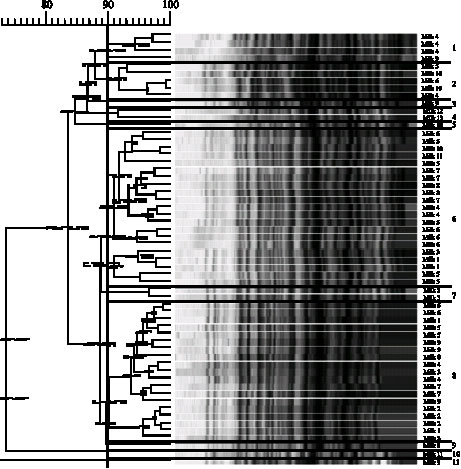
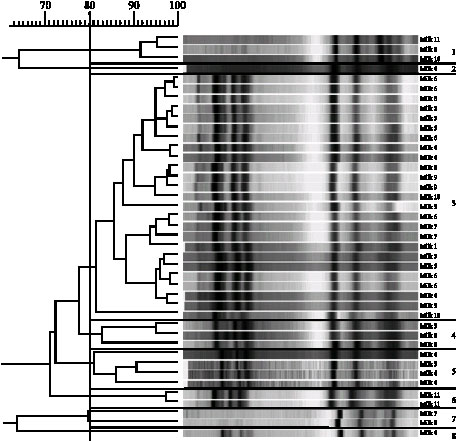
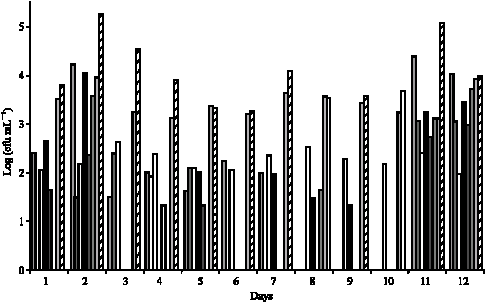
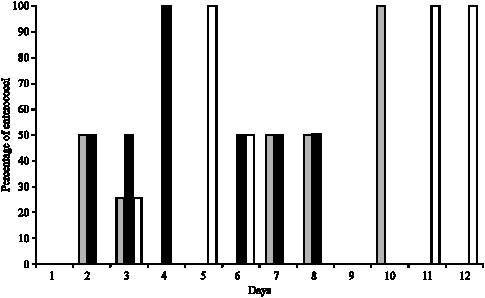
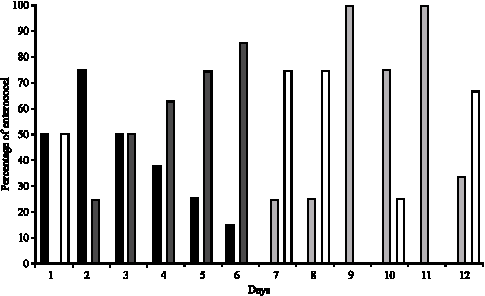
Evolution of the Raw Cow Milk Microflora, Especially Lactococci,Enterococci, Leuconostocs and Lactobacilli over a Successive 12 Day Milking Regime
Laboratoire de Microbiologie Alimentaire, ISARA-Lyon, Agrapole, 23, rue Jean Baldassini, 69364 Lyon Cedex 07, France
Prestoz Sylvie
Laboratoire de Microbiologie Alimentaire, ISARA-Lyon, Agrapole, 23, rue Jean Baldassini, 69364 Lyon Cedex 07, France
Rigobello Veronique
Laboratoire de Microbiologie Alimentaire, ISARA-Lyon, Agrapole, 23, rue Jean Baldassini, 69364 Lyon Cedex 07, France
Demarigny Yann
Laboratoire de Microbiologie Alimentaire, ISARA-Lyon, Agrapole, 23, rue Jean Baldassini, 69364 Lyon Cedex 07, France