Research Article
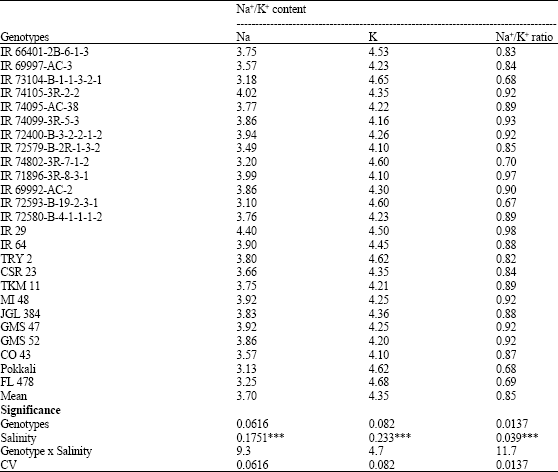
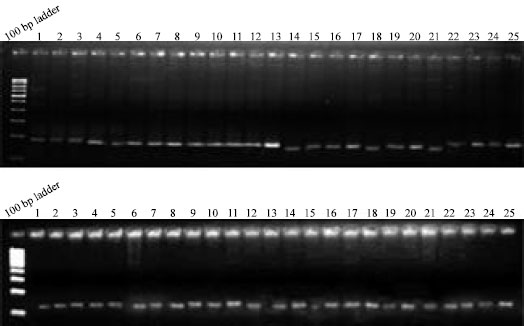
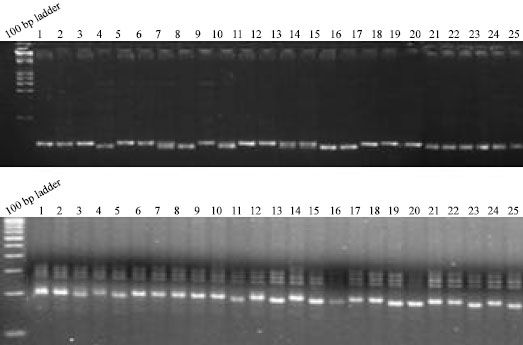
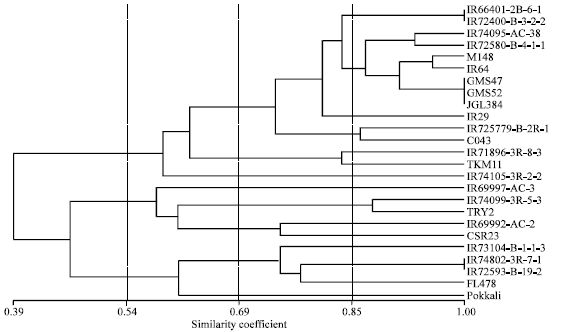
Molecular Mechanism of Salt Tolerance for Genetic Diversity Analysed in Association with Na+/K+ Ratio through SSR Markers in Rice (Oryza sativa L.)
Centre for Plant Breeding and Genetics, TNAU, Coimbatore, 641 003, India
M. Raveendran
Centre for Plant Molecular Biology, TNAU, Coimbatore, 641 003, India
C. Vijayalakshmi
Department of Plant Physiology, TNAU, Coimbatore, 641 003, India
K. Thiyagarajan
Centre for Plant Breeding and Genetics, TNAU, Coimbatore, 641 003, India
J.R. Kannan Bapu
Centre for Plant Breeding and Genetics, TNAU, Coimbatore, 641 003, India
B. C. Viraktamath
Directorate of Rice Research, Rajendranagar, Hyderabad, 500 030, India