Research Article
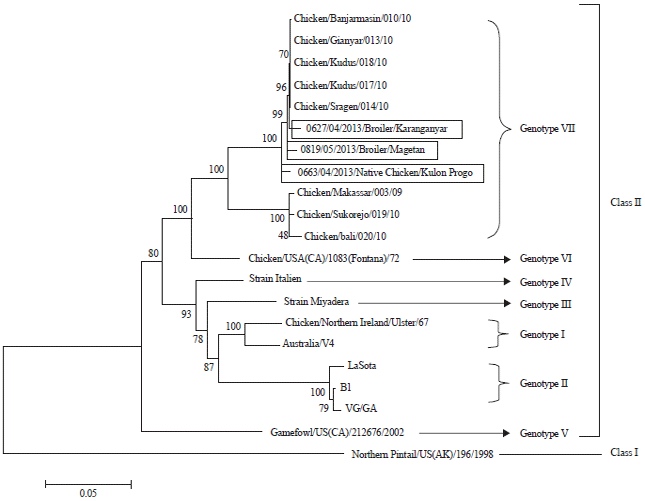
Phylogenetic Analysis of Newcastle Disease Virus from Indonesian Isolates Based on DNA-Sequence of Fusion Protein-Encoding Gene
Department of Pharmacy, Faculty of Medicine and Health Sciences, Atma Jaya Catholic University of Indonesia, Jl. Pluit Raya 2, 14440 Jakarta, Indonesia
LiveDNA: 62.23153
Nastiti Wijayanti
Department of Animal Physiology, Faculty of Biology, Universitas Gadjah Mada, Jl. Teknika Selatan, Sekip Utara, Bulaksumur, 55281 Yogyakarta, Indonesia
Aris Haryanto
Department of Biochemistry and Molecular Biology, Faculty of Veterinary Medicine, Universitas Gadjah Mada, Jl. Fauna 2, Karangmalang, 55281 Yogyakarta, Indonesia
LiveDNA: 62.12752