Research Article
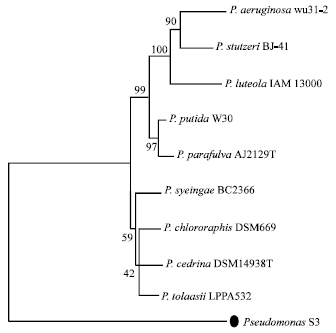
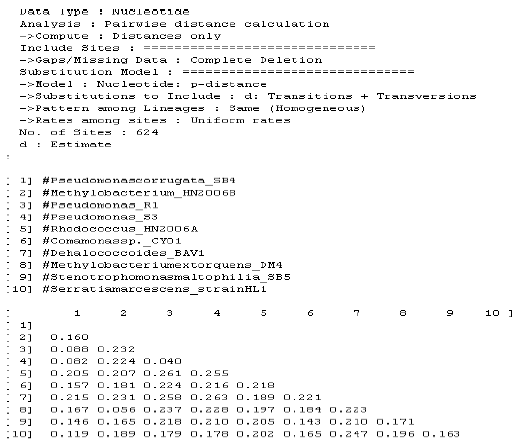
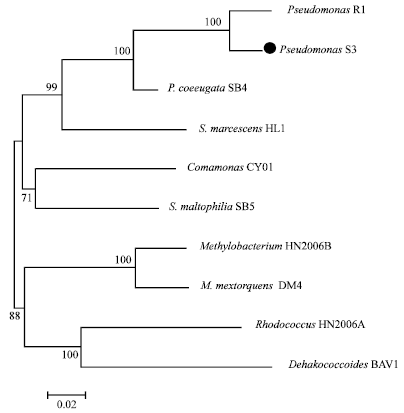
Identification of Pseudomonas sp. Strain S3 Based on Small Subunit Ribosomal RNA Gene Sequences
Department of Industrial Biotechnology, Faculty of Biosciences and Bioengineering, University Technology Malaysia, 81310 Skudai, Johor, Malaysia
S. Hamdan
Department of Industrial Biotechnology, Faculty of Biosciences and Bioengineering, University Technology Malaysia, 81310 Skudai, Johor, Malaysia
Rolando V. Pakingking Jr.
Department of Aquaculture, Southeast Asian Fisheries Development Center, Tigbauan, Iloilo, 5021, Philippines
F. Huyop
Department of Industrial Biotechnology, Faculty of Biosciences and Bioengineering, University Technology Malaysia, 81310 Skudai, Johor, Malaysia
LiveDNA: 60.18518