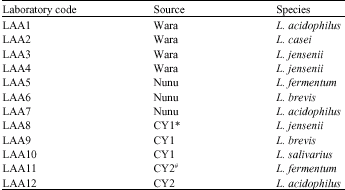
Research Article
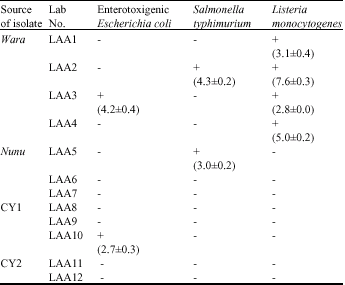
Antagonistic Effects on Enteropathogens and Plasmid Analysis of Lactobacilli Isolated from Fermented Dairy Products
Department of Biochemistry, College of Medicine, University of Lagos, Nigeria
O.R. Ejide
Department of Biochemistry, College of Medicine, University of Lagos, Nigeria
E.A. Omonigbehin
Genetics Division, Nigerian Institute for Medical Research, Yaba, Lagos, Nigeria