Research Article
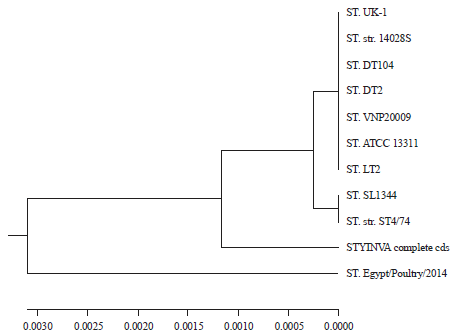
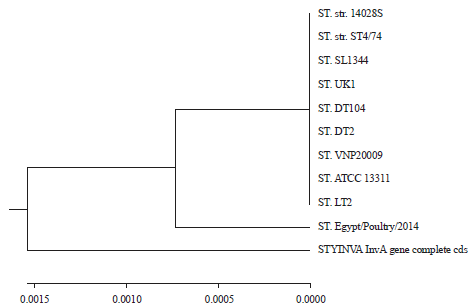
InvA Gene Sequencing of Salmonella typhimurium Isolated from Egyptian Poultry
Veterinary Serum and Vaccine Research Institute (VSVRI), Cairo, Egypt
LiveDNA: 20.15401
Hala Mohamed Abu Shady
Department of Microbiology, Faculty of Science, Ain Shams University, Cairo, Egypt
Seham Abd El-Rashed El-Zeedy
Veterinary Serum and Vaccine Research Institute (VSVRI), Cairo, Egypt
A.A. Samy
Department of Microbiology and Immunology, Veterinary Division, National Research Center (NRC), Dokki, Egypt