Research Article
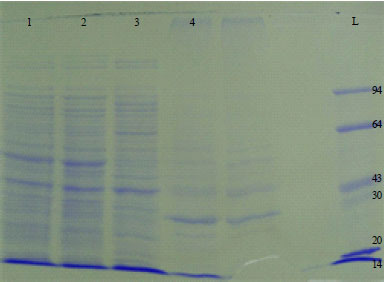
Whole Cell Protein Profiling of Human Pathogenic Bacteria Isolated from Clinical Samples
P.G and Research Department of Zoology, The New College, Chennai, Tamil Nadu, India
M. Asrar Sheriff
P.G and Research Department of Zoology, The New College, Chennai, Tamil Nadu, India