Research Article
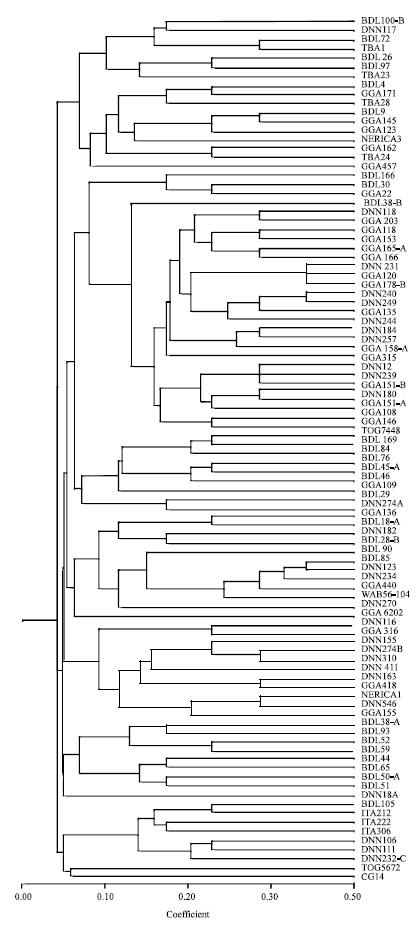
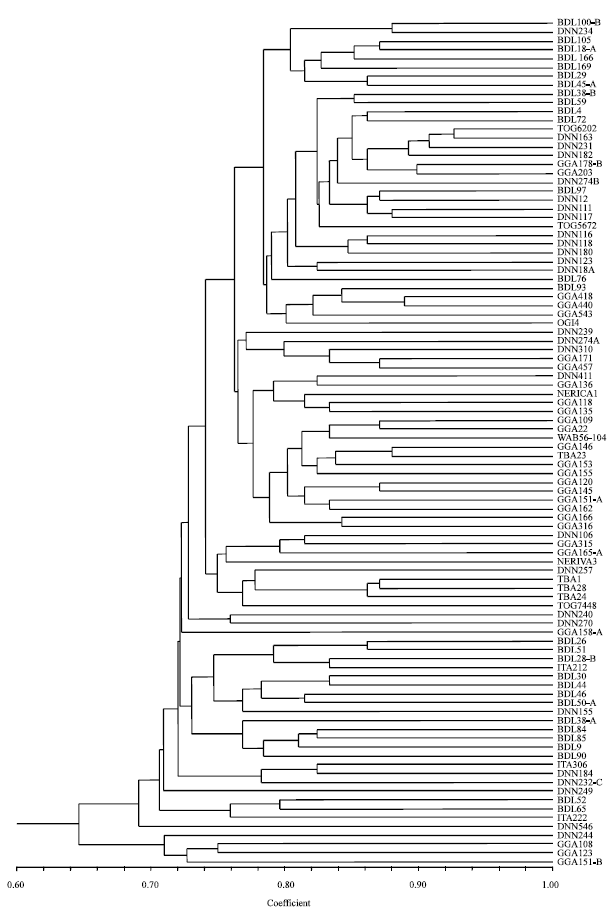
Genetic Comparisons of Landrace Rice Accessions by Morphological and RAPDs Techniques
Africa Rice Center, 01 BP 2031, Cotonou, Benin Republic
D.K. Ojo
Department of Plant Breeding and Seed Technology, University of Agriculture, Abeokuta, PMB 2240, Nigeria
A.R. Popoola
Department of Plant Breeding and Seed Technology, University of Agriculture, Abeokuta, PMB 2240, Nigeria
O.J. Ariyo
Department of Plant Breeding and Seed Technology, University of Agriculture, Abeokuta, PMB 2240, Nigeria
M. Sie
Africa Rice Center, 01 BP 2031, Cotonou, Benin Republic
K.A. Sanni
Africa Rice Center, 01 BP 2031, Cotonou, Benin Republic
F.E. Nwilene
Africa Rice Center, 01 BP 2031, Cotonou, Benin Republic
E.A. Somado
Africa Rice Center, 01 BP 2031, Cotonou, Benin Republic
R.G. Guei
FAO Viale delle Terme di Caracalla, 00100 Rome, Italy
D.D. Tia
Africa Rice Center, 01 BP 2031, Cotonou, Benin Republic
O.O. Oyelakin
International Institute of Tropical Agriculture (IITA), PMB 5320, Ibadan, Nigeria
A. Shittu
Africa Rice Center, 01 BP 2031, Cotonou, Benin Republic