Research Article
Genetic Divergence Studies in Some Indigenous Scented Rice (Oryza sativa L.) Accessions of Central India
Not Available
Abhinav Sao
Not Available
A.K. Sarawgi
Not Available
Pushpendra Singh
Not Available
Rice (Oryza sativa L.) is the main staple food crop of Asia and is being consumed by more than half of the population of the developing countries in the world. It provides 35-37% of the calories consumed by more than 3 billion Asians. It is an important food crop of the world both in terms of area (147 million ha) and production (525 million tons). In India, rice contributes around 45% of cereal production and is the main food source for more than 60% of population in the country (Siddiq, 2002).
Genetic diversity is pre-requisite for any crop improvement programme, as it helps in the development of superior recombinants (Manonmani and Fazlullah Khan, 2003). Genetic divergence among the genotypes plays an important role in selection of parents having wider variability for different characters (Nayak et al., 2004). Genetic diversity can be evaluated with morphological traits, seed proteins, isozymes and DNA markers. Conventionally it is estimated by the D2 analysis, metroglyph and principle component analysis using morphological traits (Manonmani and Fazlullah Khan, 2003). Genetic divergence analysis quantifies the genetical distance among the selected genotypes and reflects the relative contribution of specific traits towards the total divergence (Iftekharuddaula et al., 2002). The crosses between parents with maximum genetic divergence are generally the most responsive for genetic improvement (Arunachalam, 1981).
Central India is well known for its native wealth of rice genetic resources and among these the large number of indigenous short grained, scented varieties cultivated in different pockets of the Madhya Pradesh and Chhattisgarh states. These varieties in general are tall and photoperiod sensitive with aromatic, short and medium grains. Despite of low yield, they possess valuable genes for aroma and excellent cooking quality traits and enjoy emmense consumer preference within and outside the state. Therefore, genetic improvement in the yield potential of these scented accessions is needed through crossing programmes.
Thus, keeping in view the above facts, present study was conducted to estimate the nature and magnitude of genetic divergence and characters contributing to the genetic divergence were studied in fifty selected traditional aromatic rice accessions alongwith improved aromatic varieties as checks viz., Pusa Basmati, Taraori Basmati, Indira 9, Dubraj and Madhuri 11 to assess the genetic diversity among these genotypes. This study will help in selection of more distantly related parents for crossing programme to develop high yielding scented rice varieties.
The experimental material consisted of 45 scented local rice genotypes along with 5 scented varieties of rice (Table 1), collected from different districts of Chhattisgarh and Madhya Pradesh. The experiment was conducted in a Randomized Block Design with two replications at Research Farm, Department of Plant Breeding and Genetics, Indira Gandhi Agricultural University, Raipur (C.G.) during wet season, 2001. Twenty five days old seedlings were transplanted with a spacing of 20x15 cm between rows and between plants, respectively.
| Table 1: | List of used scented rice genotypes |
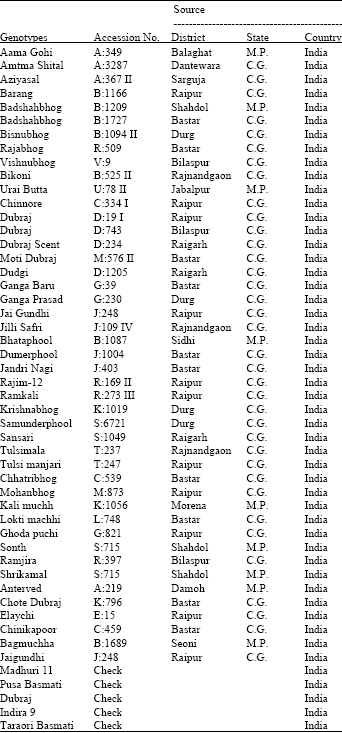 | |
| C.G. -Chhattisgarh; M.P. -Madhya Pradesh | |
Observations were recorded on five plants for 11 morphological and quality characters viz., Plant height, panicle length, Effective tillers per plant, biological yield per plant, paddy length, paddy breadth, paddy L:B (Length: Breadth) ratio, kernel length, kernel breadth, kernel L:B ratio and grain yield per plant. The mean values over replications were used for statistical analysis. The analysis of genetic divergence using Mahalanobis D2 statistics was done as described by Rao (1952) and grouping of genotypes into a number of clusters as per the standard procedure described by Spark (1973).
A clear understanding of the extent of variability prevails for each trait in germplasm would simply the job for improvement of characters through selection. But in hybridization programme where selection of genetically diverse parent is important to set wide array of recombinants, the knowledge of genetic diversity among the accession of germplasm is necessary. The Wilks test indicated highly significant differences among 50 genotypes for all the 11 characters.
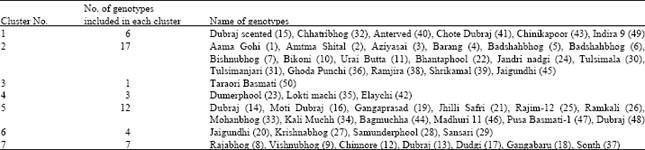
The 50 genotypes were grouped into seven clusters by using D2 values in such a way that the genotypes within a cluster had small or low D2 values than those of in between the characters. The composition of clusters has been presented in Table 2.
The maximum inter cluster distance was observed in between 3 and 4 (10.957) followed by cluster 3 and 6 (9.625), cluster 2 and 3 (8.441) and cluster 3 and 7 (7.845), whereas, minimum distance was observed in between cluster 1 and cluster 5 (2.541). The inter cluster distance varied from 2.541 to 10.957 (Table 3).
The maximum intra-cluster distance was observed for cluster 3 (2.160) followed by cluster 4 (2.051), cluster 7 (1.855) and cluster 2 (1.830) whereas, cluster 6 (1.294) observed the minimum cluster distance. Maximum number of 17 genotype were accommodated in cluster 2 followed by 12 in cluster 5, 7 in cluster 7, 6 in cluster 1 and 4 in cluster 6 and 3 in cluster 4. The minimum number of genotype 1 included in cluster 3 (Table 3).
The cluster mean for plant height was highest for cluster 6 (154.42) and lowest for cluster 3 (124.30). For effective tillers per plant cluster mean was highest to cluster 6 (8.66) while lowest for cluster 4 (4.47). The cluster mean for panicle length was highest for cluster 4 (27.93) and lowest for cluster 5 (22.24). Cluster 6 (48.51) had the highest cluster mean values biological yield per plant whereas, cluster 3 (26.41) exhibited lowest value.
| Table 2: | Genotypes included in different clusters |
 | |
| Table 3: | Average intra and inter cluster D2 values |
 | |
| Note: Intra-cluster D2- Bold and Diagonal values; Inter values are inter-cluster D2 values | |
| Table 4: | Cluster mean for morphological and quality traits |
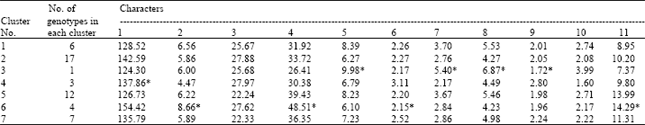 | |
| *Significance at 0.05 level, 1. Plant height (cm) 2. Effective tillers/plant 3. Panicle length (cm) 4. Biological yield/plant (g), 5. Paddy length (mm) 6. Paddy breadth (mm) 7. Paddy L:B ratio 8. Kernel length (mm), 9. Kernel breadth (mm) 10. Kernel L:B ratio 11. Grain yield/plant (g) | |
| Table 5: | Desirable genotypes for different traits |
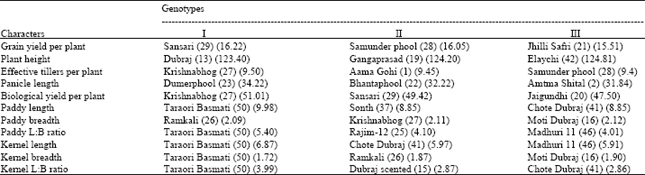 | |
| L:B ratio – Length: Breadth ratio | |
The highest cluster mean for paddy length was exhibited by cluster 3 (9.98) and lowest cluster mean value was observed for cluster 6 (6.10) (Table 4).
For quality characters, highest mean value recorded for paddy breadth was cluster 4 (3.11) and lowest for the cluster 6 (2.15) whereas, for L: B ratio of paddy highest cluster mean was shown by cluster 3 (5.40) and lowest for cluster 4 (2.17). The highest cluster mean for kernel length was shown by cluster 3 (6.87) and lowest value recorded for cluster 6 (4.23). For kernel L: B ratio the highest cluster mean value was recorded by the cluster 3 (3.99) and lowest by cluster 4 (1.60).The highest cluster mean value for grain yield per plant was exhibited by cluster 6 (14.29) and lowest by cluster 3 (7.37).
The relative divergence of each cluster from other cluster (inter cluster distance) indicated greater divergence between cluster 3 and cluster 4 followed by cluster 3 and 6, cluster 2 and cluster 3 and 3 and cluster 7 (Table 5). The selection of diverse genotype for above cluster would produce a broad spectrum of variability for morphological and quality traits studied which may enable further selection and improvement. The hybrid developed from the selected genotypes within the limits of compatibility of these cluster may produce high magnitude of heterosis or desired transgressive segregants. This would be rewarding in rice breeding programme. Sarawgi and Shrivastava (1996), Sarawgi and Rastogi (2000), Ray et al. (2002) and Nayak et al. (2004) also found similar degree of diversity in rice.
For the grain yield per plant the selected genotypes was Sansari (29), Samunderphool (28) and Jhilli Safri (21). The Genotype Dubraj (13), Gangaprasad (19) and Elaychi (42) for plant height. For effective tillers per plant the selected genotypes were Krishnabhog (27), Aama Gohi (1) and Samunderphool (28). For the panicle length the genotypes selected were Dumerphool (23) Bhataphool (22) and Amtma Shital (2).
For quality characters like Kernel length the genotypes selected were Taraori Basmati (50), Chote Dubraj (41) and Madhuri 11 (46). Chote Dubraj (41) was selected for kernel length: breadth ratio of paddy.
The above grouping indicates the existence of wide genetic divergence among constituent genotypes. Such high degree of divergence were found in local collection by Gupta et al. (1999), Sarawgi and Rastogi (2000) and Nayak et al. (2004) as well as in international collections by Usha Kumari and Rangaswamy (1997).
These observations suggest that inter crossing of genotypes from different cluster showing good mean performance may help in obtaining high yielding with good grain quality genotypes. The genotypes Tarori Basmati from cluster 3; Jaigundhi, Krishnabhog, Samunderphool and Sansari from cluster 4, Amtma Shital, Bhataphool, Ghoda Pucchi and Tulsimala from cluster 2; Dumerphool, Loktimachii and Elayachi from cluster 5 were selected. The genotypes from above different cluster can be utilized as parents in crossing programme to isolate desirable genotypes for yield and quality traits.
Sarawgi and Shrivastava (1991) reported that selection of genotypes as parents for hybridization or in crop improvement programme need not necessarily be based on geographical diversity, the genetic diversity may prove sound base for the purpose. This is in accordance to the present findings.