ABSTRACT
The study was aimed at characterizing and analyzing the genetic diversity at the isozyme level of thirty coconut cultivars. Ten enzyme systems viz., Alcohol Dehydrogenase, Glutamate Oxaloacetate Transaminase, Glucose-6 Phosphate Dehydrogenase, Peroxidase, Esterase, α-Amylase, Phosphorylase, Malate Dehydrogenase, Super Oxide Dismutase and Acid Phosphatase were studied using discontinuous polyacrylamide gel electrophoresis. Twenty loci and 40 alleles were observed, of which, 14 loci were polymorphic. Seven loci were heterozygous. Null alleles were observed for ACP, ADH and MDH. Greater heterozygosity was observed for Glucose-6 Phosphate Dehydrogenase and least for Super Oxide Dismutase. Among the cultivars, Nadora Tall and Calangute Tall showed maximum heterozygosity while Kulasekharam Green Dwarf showed least heterozygosity.
PDF Abstract XML References Citation
How to cite this article
DOI: 10.3923/ajps.2005.678.683
URL: https://scialert.net/abstract/?doi=ajps.2005.678.683
INTRODUCTION
Coconut (Cocos nucifera L.) is one of the major oilseeds of India. Out of the two types of coconut palms, Tall form is the commercially grown type and each individual is a unique heterozygous genotype than the Dwarfs. India is one of the leading countries in area as well as production of coconut in the world. However, the annual average productivity of coconut palm is low in comparison to the biological yield potential of this palm. Since, a main requirement in crop improvement programme is knowledge of the available genetic variability in the crop, collection and conservation of germplasm is of utmost importance.
In coconut, a number of cultivars are available in the different coconut growing areas of the world. At the Central Plantation Crops Research Institute (CPCRI) in Kerala, India germplasm is being collected, conserved and characterized based on morphological characters and other quantitative traits and also besides on molecular[1] and biochemical markers[2]. Additional tools for characterization are DNA fingerprinting and isozyme profiling. Isozymes are accepted as the best descriptors of genetic variability since they are codominant and also exhibit very low levels of environmental interactions[3]. This codominance also allows heterozygotes to be distinguished from homozygotes, an advantage shared by a few morphological markers. Isozymes rarely exhibit epistatic interactions, so that theoretically a genetic stock containing an infinite number of markers could be constructed[4]. Isozyme studies would help to elucidate the genetical structure of coconut populations, which would assist in adjusting the sampling strategies and would aid in screening valuable genotypes for breeding purposes. Canto-Canche et al.[5] characterized Malayan Dwarfs and Atlantic Talls by protein and enzyme electrophoresis using inflorescence and leaf extracts. Meunier[6] studied 17 enzyme systems using different tissues of the coconut palm and observed clear and consistent banding in pollen and haustorium tissues. Fernando and Gajanayake[7] used leaf tissue for isozyme analysis to characterize coconut populations used for breeding purposes. Very few workers like Geethalakshmi et al.[2] have attempted to characterize the isozyme diversity in coconut and elucidate the genetic control of the different enzyme systems that too only in dwarf cultivars. Therefore in the present study an attempt has been made to study the genetic diversity at the isozyme level using different Talls, Dwarfs and hybrids and elucidate the loci governing them and also study the level of heterozygosity.
MATERIALS AND METHODS
The studies were carried out at the division of crop improvement in the Central Plantation Crops Research Institute, Kerala, India during 1998 to 2001.
Plant material: Three palms each of thirty different coconut cultivars and hybrids viz., Fiji Tall (FJT), Ayiramkachi Tall (AYRT), Strait Settlement Green Tall (SSGT), Strait Settlement Apricot Tall (SSAT), CochinChina Tall (CCNT), Federated Malay States Tall (FMST) Andaman Giant Tall (AGT), Java Giant Tall (JVGT), Gonthembili Tall (GTBT), Seychelles Tall (SCT), Ceylon Tall (SLT), New Guinea Tall (NGAT), Nadora Tall (NDRT), Zanzibar Tall (ZAT), Standard Kudat Tall (STKT), Ganga Pani Tall (GPNT), Benaulim Tall (BENT), Calangute Tall (CALT), Spicata Tall (SPIT), Malayan Green Dwarf (MGD), King Coconut (RTB04), Kulasekharam Orange Dwarf (KOD), Kulasekharam Green Dwarf (KGD), Cameroon Red Dwarf (CRD), Kulasekharam Yellow Dwarf (KYD), Kenthali Dwarf (KTOD), Chowghat Orange Dwarf x Chowghat Orange Dwarf (COD x COD), West Coast Tall x West Coast Tall (WCT x WCT), West Coast Tall x Chowghat Orange Dwarf (WCT x COD) and Chowghat Orange Dwarf x West Coast Tall (COD x WCT) were pooled and used for isozyme analysis. Spindle leaf tissue of adult palm was found to be the best tissue for enzyme extraction.
Each sample was repeated at least twice to check the consistency of the isozyme banding pattern. Isozyme banding pattern was observed to be stable except for the minor bands of PPO and PRX and only the major bands were used for scoring and analysis.
Extraction and electrophoresis: The extraction buffer consisted of 0.1 M Tris-HCl (pH 6.8) with 10% glycerol and 0.1% β-Mercaptoethanol. A gram of leaf sample was powdered using liquid Nitrogen and extracted with 3.5 mL of extraction buffer containing 0.25 g Poly Vinyl Polypyrrolidone (PVPP). The extract was centrifuged at 10,000 g for 20 min at 4°C. The clear extract was used for enzyme electrophoresis. About 60 μL supernatant was mixed with 1.5 μL 0.01% Bromophenol Blue and loaded into the well with a micropipette or a syringe. Gels were run for 4-5 h at a constant current (25 mA/gel). Polyacrylamide gels of 7.5-12% were used to study the 10 isozyme systems-Esterase (EST), Peroxidase (PRX), Malate Dehydrogenase (MDH), Alcohol Dehydrogenase (ADH), Glutamate Oxaloacetate Transaminase (GOT), α-Amylase (α-AMY), Phosphorylase (PHOS), Glucose 6 Phosphate Dehydrogenase (G-6PDH), Super Oxide Dismutase (SOD), Acid Phosphatase (ACP) and proteins (PROT) using the standard protocols[8].
Analysis: The number of bands and loci for different isozymes were calculated for different cultivars. By noticing the pattern of bands, it was concluded whether the particular locus is homozygous or heterozygous[9].
RESULTS AND DISCUSSION
Differences in banding patterns among cultivars were observed for the different enzyme systems.
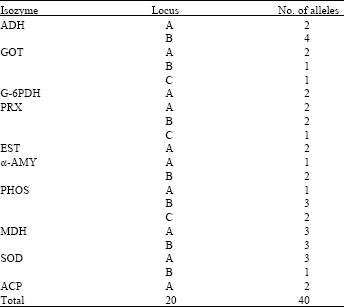
| Table 1: | Summary of genetic studies on isozyme systems |
 | |
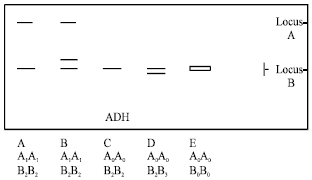 | |
| Fig. 1: | Diagrammatic representation of isozyme phenotypes of ADH |
Repeated sampling of the leaf tissue invariably gave the same set of isozyme phenotypes. The isozyme banding patterns in the systems used appear to be independent of the season, since periodic sampling of leaf tissue during the year gave consistent isozyme phenotypes. Twenty loci and 40 alleles were observed for the 10 isozyme systems, of which, 14 loci were polymorphic (70%), which is the measure of genetic variation (Table 1).
Alcohol Dehydrogenase (ADH): ADH showed five phenotypes (Fig. 1). Phenotype D showed highest frequency (Table 2). Two loci, A and B were both polymorphic and were controlled by two and four alleles, respectively. Locus B was heterozygous. KYD and KTOD did not show ADH activity possibly due to the presence of null alleles.
Glutamate Oxaloacetate Transaminase (GOT): This enzyme showed three zones of activity. Two phenotypes were observed. Phenotype B showed highest frequency (Table 2).
| Table 2: | Frequency of different isozyme phenotypes in coconut cultivars |
 | |
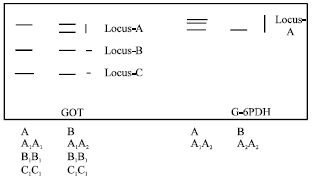 | |
| Fig. 2: | Diagrammatic representation of isozyme phenotypes of GOT and G-6PDH |
Between the three loci found to be governing this system, locus A was controlled by two alleles while loci B and C were controlled by a single allele (Fig. 2). Locus A was heterozygous.
Glucose-6 Phosphate Dehydrogenase (G-6PDH): It showed two phenotypes and of 30 cultivars studied, 28 cultivars showed phenotype A (Table 2). This enzyme is controlled by a single polymorphic locus with two alleles (Fig. 2).
Peroxidase (PRX): Six phenotypes were observed for PRX (Fig. 3). Phenotype E was seen in majority of the cultivars (Table 2). Three loci were observed with only locus A being heterozygous. Loci A and B were controlled by two alleles while locus C was controlled by one allele. Except locus C rest of the loci were polymorphic. In the present study, faint minor PRX bands were not considered for the analysis, as they were inconsistent, suggesting that their expression is influenced by external factors and/or the physiological stage of the plant. Canto-Canche et al.[5] have also observed similar inconsistencies in PRX banding pattern.
Esterase (EST): Two phenotypes were observed. Most of the cultivars expressed phenotype A (Table 2). EST was controlled by two alleles and a single polymorphic locus showing heterozygosity (Fig. 4).
α-Amylase (α-AMY): Two phenotypes were seen, of which, phenotype B was seen in majority of cultivars (Table 2). This was controlled by two loci. Locus A had one allele and locus B had two alleles (Fig. 4).
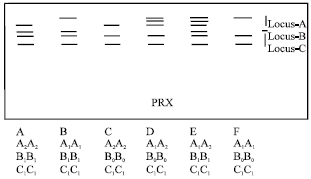 | |
| Fig. 3: | Diagrammatic representation of isozyme phenotypes of PRX |
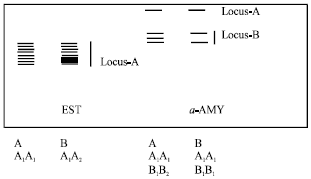 | |
| Fig. 4: | Diagrammatic representation of isozyme phenotypes of EST and α-AMY |
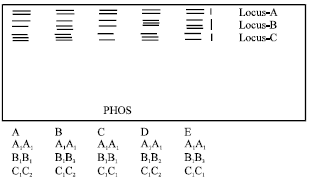 | |
| Fig. 5: | Diagrammatic representation of isozyme phenotypes of PHOS |
Phosphorylase (PHOS): Three loci and six alleles were seen for PHOS and five phenotypes were observed (Fig. 5). Phenotype C was seen in majority of the cultivars (Table 2). Loci B and C were seen to be heterozygous.
Malate Dehydrogenase (MDH): Two loci and six alleles were found for MDH (Fig. 6). Totally six phenotypes were seen, of which phenotype C was seen in majority of the cultivars (Table 2). Both loci exhibited heterozygosity. The cultivar KTOD did not exhibit any MDH activity due to the presence of null alleles.
Super Oxide Dismutase (SOD): Two loci and four alleles controlled SOD activity (Fig. 7). A total of six phenotypes were observed and frequency of phenotype A was higher (Table 2). Locus A was heterozygous.
Acid Phosphatase (ACP): It was controlled by two alleles (allele A and its null allele) of a single polymorphic locus (Fig. 8). Two phenotypes were observed for ACP, of which phenotype B (null phenotype) was seen in only one cultivar (AYRT) (Table 2).
In the present study, MDH, SOD, GOT, ADH, α-AMY and G-6PDH were found to be polymorphic as reported by other workers in other plant systems[9-13]. Earlier workers in coconut have also reported greater polymorphism for peroxidase[2,14,15]. In case of EST also polymorphism was detected in dwarf coconut cultivars[2] as in other plant systems[9,16] although in walnut it was found to be monomorphic[11]. In the present study, ACP was found to be under control of a single locus as in rye grass[17].
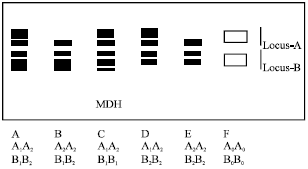 | |
| Fig. 6: | Diagrammatic representation of isozyme phenotypes of MDH |
Earlier workers have all reported monomorphism for ACP in coconut[4,17]. In oil palm, Ghesquiere[18] reported low polymorphism for ACP. The rare null allele at this locus has not been reported by any of the earlier workers. The presence of a null allele for ADH and also null alleles at the MDH loci have not been reported earlier and this could be used as an important marker in cultivar identification/breeding studies.
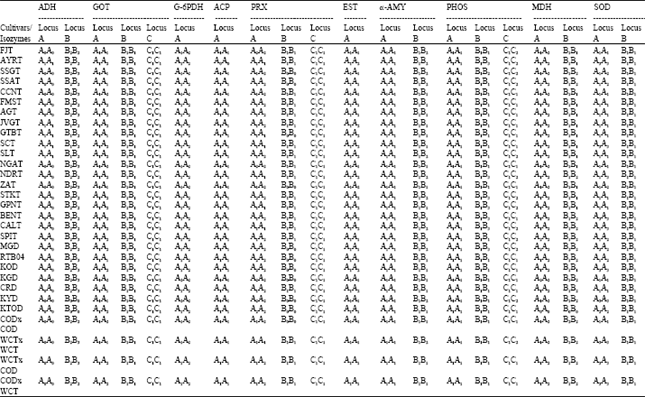
Heterozygosity: Out of the 14 loci analyzed, seven loci were heterozygous (Table 3). ADH, PRX and MDH were not used for calculating the heterozygosity because of the presence of null allele, as it was difficult to ascertain whether the band present in that locus is in homozygous or heterozygous state.
| Table 3: | Isozyme genotype for twenty polymorphic isozymes in coconut cultivars |
 | |
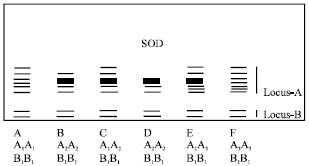 | |
| Fig. 7: | Diagrammatic representation of isozyme phenotypes of SOD |
 | |
| Fig. 8: | Diagrammatic representation of isozyme phenotypes of ACP |
Observed average heterozygosity was 26.07%. Among the seven different isozyme systems namely GOT, G-6PDH, ACP, EST, α-AMY, PHOS and SOD, highest heterozygosity (0.93) was exhibited by G-6PDH, while least heterozygosity was observed for SOD (0.23). Among the cultivars, NDRT and CALT showed maximum heterozygosity (0.54) while KGD showed least heterozygosity (0.08). Hence KGD being more homozygous can directly be used as one of the parents in breeding studies and for production of hybrids.
Heterozygotes can be identified, to a certain extent, by using isozyme profiles and these results illustrate the suitability of using isozyme profiles in identifying progeny legitimacy/parentage. Although the study of heterozygosity using isozymes is hard to find in coconut, in other plants it has been achieved[19]. In coconut, heterozygotes in both Talls and Dwarfs were observed and the number of heterozygotes in Dwarfs was found to be lower[1,20]. High heterozygosity is possibly due to the high outcrossing in the cultivars and therefore cultivars showing high heterozygosity are not suitable for use as parents in breeding studies. Such highly heterozygous cultivars need to be made more homozygous, through selection and repetitive selfing, before being used as parents in the breeding programme. High level of heterozygosity has been observed in species of olive due to the consequence of their allogamy[13].
Very few attempts have been made to characterize coconut cultivars using isozymes, even though in other genus it has been investigated[7,19,21]. An adequate genetic base for effective selection in a breeding programme is very essential. The use of isozymes as genetic marker should be of great value in coconut breeding programme where both parents are known. Isozymes are advantageous because they are expressed in young tissue as co-dominants, permitting early selections between desirable individuals (selfed or crossed progeny). Certain isozyme loci may be found to be linked to loci governing economically important morphological traits, which would allow isozyme analysis to be used to indirectly select for these morphological traits at an early stage of development, greatly increasing the efficiency of progeny evaluation by allowing coconut breeders to eliminate a portion of undesirable progeny before transplanting to the field. In the present study, SOD, PHOS, PRX and ADH showed more number of phenotypes and are useful in the characterization of cultivars. The occurrence of null alleles at the ACP, ADH and MDH loci can also be used as genetic markers in genetic studies. The above study also reveals that by using isozymes as genetic marker, breeders can improve the crop variety and can increase the palm yield potential.
ACKNOWLEDGMENTS
The authors are grateful to Indian Council of Agricultural Research for the financial assistance.
REFERENCES
- Upadhyay, A., K. Jayadev, R. Manimekalai and V.A. Parthasarathy, 2004. Genetic relationship and diversity in Indian coconut accessions based on RAPD markers. Sci. Hortic., 99: 353-362.
Direct Link - Trujillo, I., L. Rallo and A. Arus, 1995. Identifying olive cultivars by isozyme analysis. J. Am. Soc. Hortic. Sci., 120: 318-324.
Direct Link - Perera, L., J.R. Russell, J. Provan and W. Powell, 2000. Use of microsatellite DNA markers to investigate the level of genetic diversity and population genetic structure of coconut (Cocos nucifera L.). Genome, 43: 15-21.
CrossRefDirect Link








