Research Article
Structure and Molecular Evolution of Monocots Actin Genes: A cDNA Sequence and Phylogenetic Analysis of Asparagus Actin
Not Available
Toshiyuki Matsui
Not Available
Haruo Suzuki
Not Available
Yusuke Kosugi
Not Available
The different cellular processes in which actin plays demonstrated or proposed roles include establishing cell polarity, division plane determination (by positioning the prepophase band), preprogramming of development and cell wall deposition, cell elongation, tip growth (e.g., of pollen tubes, root hairs and moss protonema), transmembrane transport and positioning of receptors, mRNA transport within the cell, cytoplasmic streaming and orientation of chloroplasts in response of light with appropriate repositioning of the nucleus (Staiger and Lloyd, 1991; Staehelin and Hepler, 1996). Because plant cells do not migrate, establishment of polarity, division plane determination, cell elongation and directing cell wall deposition are processes critical to nearly all aspects of plant development and morphology.
With the recent explosion of information in plant genomics, much is already known about the evolution of actins in mammals, invertebrates, insects and other representatives of major linages (Drouin and Moniz, 1996). In monocots, however, only a few representatives of actin multigene family have been cloned and sequenced and little is known about their pattern and process of evolution. In angiosperms, actin is encoded by a relatively large, diverse and dispersed multigene family comprising 8-40 genes (Bernatzky and Tanksley, 1986). As in most other eukaryotes, the multiple copies of actin genes in angiosperms are thought to allow a more diverse pattern of gene regulation, rather than direct the production of a large amount of actin (Hightower and Meagher, 1986). Based on the shared intron positions and the many short amino acid sequences unique to angiosperm sequences, it is believed that all angiosperm actin sequences are derived from a single ancestral sequence. Relative to other nuclear genes, actin genes are highly conserved in both size and sequence (Li and Graur, 1991). However, relative to mammalian actins, angiosperm actin genes show a greater sequence divergence. It has been suggested that angiosperm actin are evolving at the same rate as those in fungi and mammals and that their relatively high sequence divergence is due to a more ancient origin.
This study focuses on the sequencing and sequence data analysis of an asparagus actin gene to improve our understanding about structure of actin gene in asparagus. Phylogenetic analysis has been used to gain some understanding of the pattern of actin gene evolution.
RNA isolation and PCR amplification: Total RNA was extracted from asparagus (Asparagus officinnalis L. cv. Welcome) spears using Hot Borate method adapted from Wan and Wilkins (1994). The first strand cDNA was synthesized from 2 g of the total RNA by reverse transcriptase with Oligo-(dT) primer according to the instruction of SUPER SCRIPTTM Preamplification System for First Strand cDNA Synthesis (GIBCOBRL, Tokyo, Japan). PCR was performed in a total volume of 25μl containing the first strand cDNA reaction products, 10xPCR Buffer, MgCl2, dNTP, First Start Taq DNA Polymerase (Roche) and primers. The primers (5’-GARAARATGACNCARATHATG-3’ as the upstream primer and 5’-TCNACRTCRCAYTTCATDAT -3’ as the downstream primer) were designed and synthesized on the basis of amino acid domains (EKMTQIM and IMKCDVD respectively) conserved in various actin genes. The Sal 1 and Not 1 restriction site sequences were also included at 5’-end of the sense and antisense primer, to facilitate cloning of PCR product. The PCR procedure started with 10 min at 950C and was carried out 35 cycles of 30 s at 950C, 30 s at 500C and 30 s at 720C and 10 min at 720C with ASTEC Program Temperature Control System PC-700. The PCR products were confirmed by agarose gel electrophoresis.
Cloning and sequencing of cDNA: The amplified cDNA was ligated to the plasmid pSPORT1 and cloned into Escherichia coli (DH-5α ) Not 1- Sal 1 – cut (BRL, Tokyo, Japan). Sequencing was performed by the cycle sequencing method using GATCR-Bio Cycle sequencing Kit and a DNA sequencer GATC 1500 Long-Run system (GATC Gmbh, Konstanz, Germany).
Sequence data analysis: Sequence analysis was performed using computer software GENETYX-MAC Ver.7. Homology searches with the Genbank and the EMBL databases were performed using the homology program in the software. The phylogenetic tree was also constructed with the UPGMA method in the software.
Results and Discussions
PCR amplification of actin gene: The amino and carboxyl termini of plant actins are conserved. Based on this fact, degenerate primers were designed using the plant actin sequences already available. The cDNA pAS-AC is a partial clone encoding actin in asparagus. The sense primer hybridizes to the region from the first nucleotide of EKMTQIM and the antisense primer hybridizes to the IMKCDVD.
Variation in amino acid and nucleotide sequences: The encoded mRNA of pAS-AC is 515 bp long (Fig. 1) and is highly homologus to actin genes of other plants.
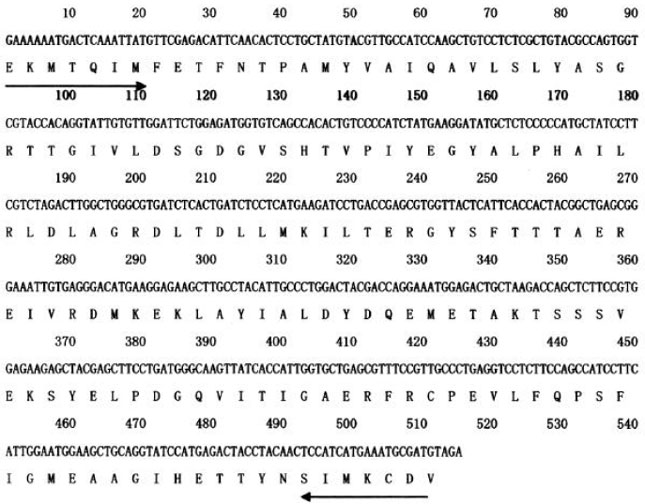 | |
| Fig. 1: | Necleotide sequence and deduced amino acid sequence of the cDNA clone corresponding to pAS-AC. The predicted amino acid sequence is given in single-letter code for each amino acid. The arrows indicate the positions of degenerated primers (→ sense, ← antisense) used for RT-PCR. Numbering refers to total nucleotide residues on each line |
 | |
| Fig. 2: | Comparison of the deduced amino acid sequences from asparagus (AB111527), banana (AB022041), maize (U60508), rice (X15865) and sorghum (X79378) by multi alignment. The amino acid residues are numbered at the beginning and end of the sequences on each line. Asterisks (*) denote the amino acid residues those are identical. Dashes in the amino acid sequences represent gaps introduced to maximize alignment of the polypeptides. |
| Table 1: | Percentage of nucleotide and deduced amino acid homology between actin from asparagus spear and other plants in the databases. |
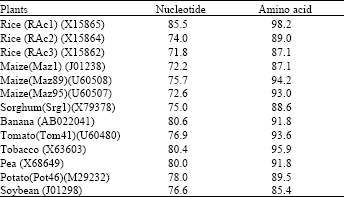 | |
| Asparagus actin (AB111527) is calculated as 100 % | |
The pAS-AC sequence is 85.5% identical to actin from rice (X15865), 80.6% identical to actin from banana (AB022041) and 75.7% to maize actin (U60508). Allowing for conservative amino acid substitutions, the similarities are 98.2, 91.8 and 87.1% for rice, banana and maize sequence respectively, Actin nucleotide sequences vary by as much as 21-25% both within and between species (Drouin and Moniz, 1996). This suggests that the actin multigene family is equally old in all lineages. The fact that this level of variation is also observed between the Volvox actin gene and all the angiosperm actin genes suggests that variation in nucleotide sequence between some angiosperm actin genes is saturated. Actin sequences in monocots vary from 0-14% at the amino acid level. Within a species, the variation in amino acid sequences is 2-12% in maize.
GC content of actin gene: The GC content of monocot actin genes used in this study varies with in a relatively narrow range. The average GC content of dicot actin genes is lower than that of monocot actin genes. Drouin and Moniz ( 1996) suggested that monocot and dicot actin genes have different codon usages patterns and the difference in GC content between monocot and dicot actin genes is solely due to a GC bias at the third positions of codons.
Phylogeny of asparagus actin gene: The asparagus actin sequenced in this study was aligned with 13 existing actin sequences from maize soybean, potato, tobacco, rice, pea, sorghum, tomato and banana. This alignment was used to generate phylogenetic tree. Nucleotide sequences, rather than amino acid sequences, were used to construct tree because they had twice the number of variable. The order of sequence input had no effect on the topology of the tree. The topology in the tree suggests that there might be two main lineages of angiosperm actin genes, one group composed exclusively of dicot actin genes and a second group composed of both monocot and dicot actin genes. Asparagus actin falls into the second main lineage.
The phylogenetic analysis of asparagus actin gene presented here showed that, despite the fact that the actin multigene family underwent several gene duplication events that preceded the monocot-dicot divergence, most of the monocot actin genes clustered together on the tree.
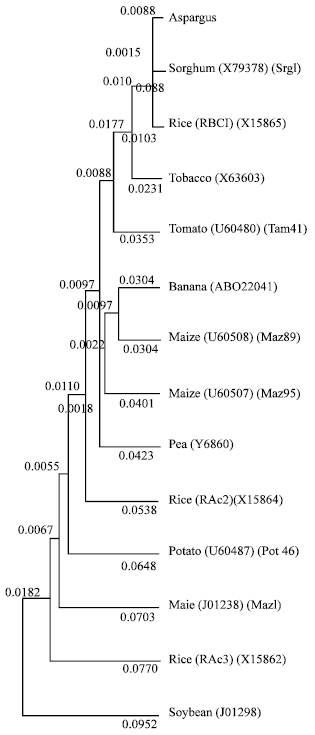 | |
| Fig. 3: | Phylogenetic tree of the alignment of pAS-AC deduced amino acid sequence with other actin gene in the database. Protein sequences were aligned using UPGMA and a phylogenetic tree was constructed using GENETYX-MAC software. The GenBank accession numbers are shown in the parentheses. |
This clustering of the monocot actin genes suggests that these sequences may have undergone concerted evolution early in the evolution of monocots. The work presented here has laid the groundwork for further characterization of the evolution and function of actin genes in asparagus.